The domain within your query sequence starts at position 708 and ends at position 737; the E-value for the ANK domain shown below is 3.95e1.
LKNTMLERACDQNNSIMVECLLLLGADANQ
ANKankyrin repeats |
|---|
| SMART accession number: | SM00248 |
|---|---|
| Description: | Ankyrin repeats are about 33 amino acids long and occur in at least four consecutive copies. They are involved in protein-protein interactions. The core of the repeat seems to be an helix-loop-helix structure. |
| Interpro abstract (IPR002110): | The ankyrin repeat is one of the most common protein-protein interaction motifs in nature. Ankyrin repeats are tandemly repeated modules of about 33 amino acids. They occur in a large number of functionally diverse proteins mainly from eukaryotes. The few known examples from prokaryotes and viruses may be the result of horizontal gene transfers. The repeat has been found in proteins of diverse function such as transcriptional initiators, cell-cycle regulators [ (PUBMED:31000436) ], cytoskeletal, ion transporters and signal transducers [ (PUBMED:29769718) (PUBMED:8108379) ]. The ankyrin fold appears to be defined by its structure rather than its function since there is no specific sequence or structure which is universally recognised by it. The conserved fold of the ankyrin repeat unit is known from several crystal and solution structures [ (PUBMED:8875926) (PUBMED:9353127) (PUBMED:9461436) (PUBMED:9865693) ]. Each repeat folds into a helix-loop-helix structure with a beta-hairpin/loop region projecting out from the helices at a 90 o angle. The repeats stack together to form an L-shaped structure [ (PUBMED:8875926) (PUBMED:12461176) ]. |
| GO function: | protein binding (GO:0005515) |
| Family alignment: |
There are 1471345 ANK domains in 265757 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing ANK domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with ANK domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing ANK domain in the selected taxonomic class.
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Mandiyan V, Andreev J, Schlessinger J, Hubbard SR
- Crystal structure of the ARF-GAP domain and ankyrin repeats of PYK2-associated protein beta.
- EMBO J. 1999; 18: 6890-8
- Display abstract
ADP ribosylation factors (ARFs), which are members of the Ras superfamily of GTP-binding proteins, are critical components of vesicular trafficking pathways in eukaryotes. Like Ras, ARFs are active in their GTP-bound form, and their duration of activity is controlled by GTPase-activating proteins (GAPs), which assist ARFs in hydrolyzing GTP to GDP. PAPbeta, a protein that binds to and is phosphorylated by the non-receptor tyrosine kinase PYK2, contains several modular signaling domains including a pleckstrin homology domain, an SH3 domain, ankyrin repeats and an ARF-GAP domain. Sequences of ARF-GAP domains show no recognizable similarity to those of other GAPs, and contain a characteristic Cys-X(2)-Cys-X(16-17)-Cys-X(2)-Cys motif. The crystal structure of the PAPbeta ARF-GAP domain and the C-terminal ankyrin repeats has been determined at 2.1 A resolution. The ARF-GAP domain comprises a central three-stranded beta-sheet flanked by five alpha-helices, with a Zn(2+) ion coordinated by the four cysteines of the cysteine-rich motif. Four ankyrin repeats are also present, the first two of which form an extensive interface with the ARF-GAP domain. An invariant arginine and several nearby hydrophobic residues are solvent exposed and are predicted to be the site of interaction with ARFs. Site-directed mutagenesis of these residues confirms their importance in ARF-GAP activity.
- Jacobs MD, Harrison SC
- Structure of an IkappaBalpha/NF-kappaB complex.
- Cell. 1998; 95: 749-58
- Display abstract
The inhibitory protein, IkappaBalpha, sequesters the transcription factor, NF-kappaB, as an inactive complex in the cytoplasm. The structure of the IkappaBalpha ankyrin repeat domain, bound to a partially truncated NF-kappaB heterodimer (p50/ p65), has been determined by X-ray crystallography at 2.7 A resolution. It shows a stack of six IkappaBalpha ankyrin repeats facing the C-terminal domains of the NF-kappaB Rel homology regions. Contacts occur in discontinuous patches, suggesting a combinatorial quality for ankyrin repeat specificity. The first two repeats cover an alpha helically ordered segment containing the p65 nuclear localization signal. The position of the sixth ankyrin repeat shows that full-length IkappaBalpha will occlude the NF-kappaB DNA-binding cleft. The orientation of IkappaBalpha in the complex places its N- and C-terminal regions in appropriate locations for their known regulatory functions.
- Bork P
- Hundreds of ankyrin-like repeats in functionally diverse proteins: mobile modules that cross phyla horizontally?
- Proteins. 1993; 17: 363-74
- Display abstract
Based on pattern searches and systematic database screening, almost 650 different ankyrin-like (ANK) repeats from nearly all phyla have been identified; more than 150 of them are reported here for the first time. Their presence in functionally diverse proteins such as enzymes, toxins, and transcription factors strongly suggests domain shuffling, but their occurrence in prokaryotes and yeast excludes exon shuffling. The spreading mechanism remains unknown, but in at least three cases horizontal gene transfer appears to be involved. ANK repeats occur in at least four consecutive copies. The terminal repeats are more variable in sequence. One feature of the internal repeats is a predicted central hydrophobic alpha-helix, which is likely to interact with other repeats. The functions of the ankyrin-like repeats are compatible with a role in protein-protein interactions.
- Davis LH, Otto E, Bennett V
- Specific 33-residue repeat(s) of erythrocyte ankyrin associate with the anion exchanger.
- J Biol Chem. 1991; 266: 11163-9
- Display abstract
Erythrocyte ankyrin contains an 89-kDa domain (residues 2-827) comprised almost entirely of 22 tandem repeats of 33 amino acids which are responsible for the high affinity interaction of ankyrin with the anion exchanger (Davis, L., and Bennett, V. (1990) J. Biol. Chem. 265, 10589-10596). The question of whether the repeats are equivalent with respect to binding to the anion exchanger was addressed using defined regions of erythrocyte and brain ankyrins expressed in bacteria. The conclusion is that the repeats are not interchangeable and that the 44 residues from 722 to 765 are essential for high affinity binding between erythrocyte ankyrin and the anion exchanger. Residues 348-765 were active whereas a polypeptide of the same size (residues 305-721) but missing the 44 residues was not active. The difference between the active and inactive polypeptides was not caused by the degree of folding based on circular dichroism spectra. The 44 residues from 722 to 765 were not sufficient for binding since deletions of residues from 348 to 568 resulted in a 10-fold loss of activity. However, the role of residues 348-568 may be at the level of folding rather than a direct contact since the deleted sequences were not active in the absence of 722-765 and since circular dichroism spectra revealed significant loss of structure in the smaller polypeptides. Further evidence that the 33-residue repeats are not equivalent in ability to bind to the anion exchanger is that a region of human brain ankyrin containing 18 33-residue repeats with 67% overall sequence identity to erythrocyte ankyrin was 8-fold less active than a region of erythrocyte ankyrin containing only 12 repeats. The fact that the anion exchanger binds to certain repeats suggests that the other 33-amino acid repeats could interact with proteins distinct from the anion exchanger and provide ankyrin with the potential for considerable diversity in association with membrane proteins as well as cytoplasmic proteins. Tubulin was identified as one example of a protein that can interact with ankyrin repeats that are not recognized by the anion exchanger.
- Lux SE, John KM, Bennett V
- Analysis of cDNA for human erythrocyte ankyrin indicates a repeated structure with homology to tissue-differentiation and cell-cycle control proteins.
- Nature. 1990; 344: 36-42
- Display abstract
Analysis of complementary DNA for human erythroid ankyrin indicates that the mature protein contains 1,880 amino acids comprising an N-terminal domain binding integral membrane proteins and tubulin, a central domain binding spectrin and vimentin, and an acidic C-terminal 'regulatory' domain containing an alternatively spliced sequence missing from ankyrin variant 2.2. The N-terminal domain is almost entirely composed of 22 tandem 33-amino-acid repeats. Similar repeats are found in yeast and invertebrate proteins involved in cell-cycle control and tissue differentiation.
- Disease (disease genes where sequence variants are found in this domain)
-
SwissProt sequences and OMIM curated human diseases associated with missense mutations within the ANK domain.
Protein Disease Ankyrin-1 (P16157) (SMART) OMIM:182900: Spherocytosis-2 DNA-binding protein RFXANK (O14593) (SMART) OMIM:603200: MHC class II deficiency, complementation group B
OMIM:209920: - Metabolism (metabolic pathways involving proteins which contain this domain)
-
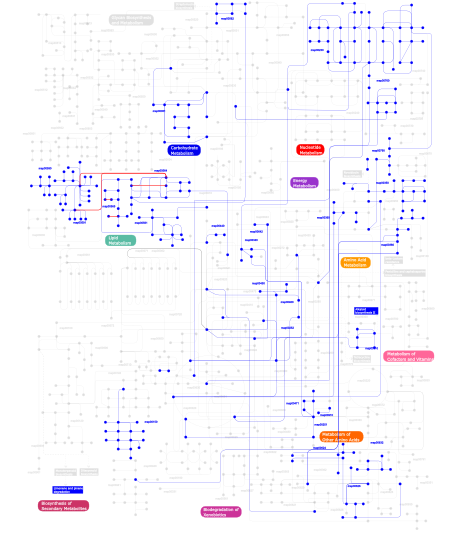
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 5.56 map04662 B cell receptor signaling pathway 5.56 map04660 T cell receptor signaling pathway 5.56 map04920 Adipocytokine signaling pathway 4.42 map05120 Epithelial cell signaling in Helicobacter pylori infection 3.99 map05222 Small cell lung cancer 3.85 map04320 Dorso-ventral axis formation 3.85 map04330 Notch signaling pathway 3.71 map05220 Chronic myeloid leukemia 3.57 map04210 Apoptosis 3.57 map04620 Toll-like receptor signaling pathway 3.57 map05215 Prostate cancer 2.85 map04010 MAPK signaling pathway 2.57  map00910
map00910Nitrogen metabolism 2.43  map00564
map00564Glycerophospholipid metabolism 2.28 map05212 Pancreatic cancer 2.28 map04510 Focal adhesion 2.14  map00471
map00471D-Glutamine and D-glutamate metabolism 2.14 map05221 Acute myeloid leukemia 2.14  map00251
map00251Glutamate metabolism 2.14 map04810 Regulation of actin cytoskeleton 2.00 map04111 Cell cycle - yeast 1.85 map04110 Cell cycle 1.85 map03050 Proteasome 1.57 map04070 Phosphatidylinositol signaling system 1.57 map05219 Bladder cancer 1.43  map00561
map00561Glycerolipid metabolism 1.43  map00310
map00310Lysine degradation 1.14 map05213 Endometrial cancer 1.14 map04720 Long-term potentiation 1.14  map00380
map00380Tryptophan metabolism 1.14 map03320 PPAR signaling pathway 1.00 map04612 Antigen processing and presentation 1.00 map05020 Parkinson's disease 0.86  map00760
map00760Nicotinate and nicotinamide metabolism 0.86 map04080 Neuroactive ligand-receptor interaction 0.71 map04664 Fc epsilon RI signaling pathway 0.71 map04730 Long-term depression 0.71 map04370 VEGF signaling pathway 0.71  map00590
map00590Arachidonic acid metabolism 0.71  map00565
map00565Ether lipid metabolism 0.71 map04912 GnRH signaling pathway 0.71 map00592 alpha-Linolenic acid metabolism 0.71  map00591
map00591Linoleic acid metabolism 0.43  map00350
map00350Tyrosine metabolism 0.43 map04350 TGF-beta signaling pathway 0.43  map00460
map00460Cyanoamino acid metabolism 0.43  map00632
map00632Benzoate degradation via CoA ligation 0.43  map00252
map00252Alanine and aspartate metabolism 0.43  map00340
map00340Histidine metabolism 0.43  map00562
map00562Inositol phosphate metabolism 0.29  map00150
map00150Androgen and estrogen metabolism 0.29  map00626
map00626Naphthalene and anthracene degradation 0.29  map00051
map00051Fructose and mannose metabolism 0.29  map00440
map00440Aminophosphonate metabolism 0.29  map00450
map00450Selenoamino acid metabolism 0.14  map00642
map00642Ethylbenzene degradation 0.14 map05214 Glioma 0.14 map04115 p53 signaling pathway 0.14  map00360
map00360Phenylalanine metabolism 0.14  map00790
map00790Folate biosynthesis 0.14 map03060 Protein export 0.14 map05223 Non-small cell lung cancer 0.14 map00903 Limonene and pinene degradation 0.14 map00960 Alkaloid biosynthesis II 0.14  map00624
map006241- and 2-Methylnaphthalene degradation 0.14  map00230
map00230Purine metabolism 0.14 map05218 Melanoma This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with ANK domain which could be assigned to a KEGG orthologous group, and not all proteins containing ANK domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of ANK domains in PDB
PDB code Main view Title 1a5e 
SOLUTION NMR STRUCTURE OF TUMOR SUPPRESSOR P16INK4A, 18 STRUCTURES 1ap7 
P19-INK4D FROM MOUSE, NMR, 20 STRUCTURES 1awc 
MOUSE GABP ALPHA/BETA DOMAIN BOUND TO DNA 1bd8 
STRUCTURE OF CDK INHIBITOR P19INK4D 1bi7 
MECHANISM OF G1 CYCLIN DEPENDENT KINASE INHIBITION FROM THE STRUCTURE OF THE CDK6-P16INK4A TUMOR SUPPRESSOR COMPLEX 1bi8 
MECHANISM OF G1 CYCLIN DEPENDENT KINASE INHIBITION FROM THE STRUCTURES CDK6-P19INK4D INHIBITOR COMPLEX 1blx 
P19INK4D/CDK6 COMPLEX 1bu9 
SOLUTION STRUCTURE OF P18-INK4C, 21 STRUCTURES 1d9s 
TUMOR SUPPRESSOR P15(INK4B) STRUCTURE BY COMPARATIVE MODELING AND NMR DATA 1dc2 
SOLUTION NMR STRUCTURE OF TUMOR SUPPRESSOR P16INK4A, 20 STRUCTURES 1dcq 
CRYSTAL STRUCTURE OF THE ARF-GAP DOMAIN AND ANKYRIN REPEATS OF PAPBETA. 1g3n 
STRUCTURE OF A P18(INK4C)-CDK6-K-CYCLIN TERNARY COMPLEX 1ihb 
CRYSTAL STRUCTURE OF P18-INK4C(INK6) 1ikn 
IKAPPABALPHA/NF-KAPPAB COMPLEX 1ixv 
Crystal Structure Analysis of homolog of oncoprotein gankyrin, an interactor of Rb and CDK4/6 1k1a 
Crystal structure of the ankyrin repeat domain of Bcl-3: a unique member of the IkappaB protein family 1k1b 
Crystal structure of the ankyrin repeat domain of Bcl-3: a unique member of the IkappaB protein family 1k3z 
X-ray crystal structure of the IkBb/NF-kB p65 homodimer complex 1mj0 
SANK E3_5: an artificial Ankyrin repeat protein 1mx2 
Structure of F71N mutant of p18INK4c 1mx4 
Structure of p18INK4c (F82Q) 1mx6 
Structure of p18INK4c (F92N) 1myo 
SOLUTION STRUCTURE OF MYOTROPHIN, NMR, 44 STRUCTURES 1n0q 
3ANK: A designed ankyrin repeat protein with three identical consensus repeats 1n0r 
4ANK: A designed ankyrin repeat protein with four identical consensus repeats 1n11 
D34 REGION OF HUMAN ANKYRIN-R AND LINKER 1nfi 
I-KAPPA-B-ALPHA/NF-KAPPA-B COMPLEX 1ot8 
Structure of the Ankyrin Domain of the Drosophila Notch Receptor 1oy3 
CRYSTAL STRUCTURE OF AN IKBBETA/NF-KB P65 HOMODIMER COMPLEX 1qym 
X-ray structure of human gankyrin 1s70 
Complex between protein ser/thr phosphatase-1 (delta) and the myosin phosphatase targeting subunit 1 (MYPT1) 1svx 
Crystal structure of a designed selected Ankyrin Repeat protein in complex with the Maltose Binding Protein 1sw6 
S. CEREVISIAE SWI6 ANKYRIN-REPEAT FRAGMENT 1tr4 
Solution structure of human oncogenic protein gankyrin 1uoh 
HUMAN GANKYRIN 1wdy 
Crystal structure of ribonuclease 1wg0 
Structural comparison of Nas6p protein structures in two different crystal forms 1ycs 
P53-53BP2 COMPLEX 1ymp 
The Crystal Structure of a Partial Mouse Notch-1 Ankyrin Domain: Repeats 4 Through 7 Preserve an Ankyrin Fold 1yyh 
Crystal structure of the human Notch 1 ankyrin domain 2a5e 
SOLUTION NMR STRUCTURE OF TUMOR SUPPRESSOR P16INK4A, RESTRAINED MINIMIZED MEAN STRUCTURE 2aja 
X-Ray structure of an ankyrin repeat family protein Q5ZSV0 from Legionella pneumophila. Northeast Structural Genomics Consortium target LgR21. 2b0o 
Crystal structure of UPLC1 GAP domain 2bkg 
Crystal structure of E3_19 an designed ankyrin repeat protein 2bkk 
Crystal structure of Aminoglycoside Phosphotransferase APH(3')-IIIa in complex with the designed ankyrin repeat inhibitor AR_3a 2di3 
Crystal structure of the transcriptional factor CGL2915 from Corynebacterium glutamicum 2dvw 
Structure of the Oncoprotein Gankyrin in Complex with S6 ATPase of the 26S Proteasome 2dwz 
Structure of the Oncoprotein Gankyrin in Complex with S6 ATPase of the 26S Proteasome 2dzn 
Crystal structure analysis of yeast Nas6p complexed with the proteasome subunit, rpt3 2dzo 
Crystal structure analysis of yeast Nas6p complexed with the proteasome subunit, rpt3 2eta 
Crystal structure of the ankyrin repeat domain of the TRPV2 2etb 
Crystal structure of the ankyrin repeat domain of TRPV2 2etc 
Crystal structure of the ankyrin repeat domain of TRPV2 2f37 
Crystal structure of the ankyrin repeat domain of human TRPV2 2f8x 
Crystal structure of activated Notch, CSL and MAML on HES-1 promoter DNA sequence 2f8y 
Crystal structure of human Notch1 ankyrin repeats to 1.55A resolution. 2fo1 
Crystal Structure of the CSL-Notch-Mastermind ternary complex bound to DNA 2he0 
Crystal structure of a human Notch1 ankyrin domain mutant 2j8s 
Drug Export Pathway of Multidrug Exporter AcrB Revealed by DARPin Inhibitors 2jab 
A designed ankyrin repeat protein evolved to picomolar affinity to Her2 2kbx 
Solution structure of ILK-PINCH complex 2kxp 
Solution NMR structure of V-1 bound to capping protein (CP) 2l6b 
NRC consensus ankyrin repeat protein solution structure 2myo 
SOLUTION STRUCTURE OF MYOTROPHIN, NMR, MINIMIZED AVERAGE STRUCTURE 2nyj 
Crystal structure of the ankyrin repeat domain of TRPV1 2p2c 
Inhibition of caspase-2 by a designed ankyrin repeat protein (DARPin) 2pnn 
Crystal Structure of the Ankyrin Repeat Domain of Trpv1 2qc9 
Mouse Notch 1 Ankyrin Repeat Intracellular Domain 2qyj 
Crystal structure of a designed full consensus ankyrin 2rfa 
Crystal structure of the mouse TRPV6 ankyrin repeat domain 2rfm 
Structure of a Thermophilic Ankyrin Repeat Protein 2v4h 
NEMO CC2-LZ domain - 1D5 DARPin complex 2v5q 
CRYSTAL STRUCTURE OF WILD-TYPE PLK-1 KINASE DOMAIN IN COMPLEX WITH A SELECTIVE DARPIN 2vge 
Crystal structure of the C-terminal region of human iASPP 2xee 
Structural Determinants for Improved Thermal Stability of Designed Ankyrin Repeat Proteins With a Redesigned C-capping Module. 2xeh 
Structural Determinants for Improved Thermal Stability of Designed Ankyrin Repeat Proteins With a Redesigned C-capping Module. 2xen 
Structural Determinants for Improved Thermal Stability of Designed Ankyrin Repeat Proteins With a Redesigned C-capping Module. 2xzd 
Caspase-3 in Complex with an Inhibitory DARPin-3.4 2xzt 
Caspase-3 in Complex with DARPin-3.4_I78S 2y0b 
Caspase-3 in Complex with an Inhibitory DARPin-3.4_S76R 2y1l 
Caspase-8 in Complex with DARPin-8.4 2zgd 
Asn-hydroxylation stabilises the ankyrin repeat domain fold 2zgg 
Asn-hydroxylation stabilises the ankyrin repeat domain fold 3aaa 
Crystal Structure of Actin capping protein in complex with V-1 3aji 
Structure of Gankyrin-S6ATPase photo-cross-linked site-specifically, and incoporated by genetic code expansion 3b7b 
EuHMT1 (Glp) Ankyrin Repeat Domain (Structure 1) 3b95 
EuHMT1 (Glp) Ankyrin Repeat Domain (Structure 2) 3c5r 
Crystal Structure of the BARD1 Ankyrin Repeat Domain and Its Functional Consequences 3d9h 
Crystal Structure of the Splice Variant of Human ASB9 (hASB9-2), an Ankyrin Repeat Protein 3deo 
Structural basis for specific substrate recognition by the chloroplast signal recognition particle protein cpSRP43 3dep 
Structural basis for specific substrate recognition by the chloroplast signal recognition particle protein cpSRP43 3ehq 
Crystal Structure of Human Osteoclast Stimulating Factor 3ehr 
Crystal Structure of Human Osteoclast Stimulating Factor 3eu9 
The ankyrin repeat domain of Huntingtin interacting protein 14 3f6q 
Crystal structure of integrin-linked kinase ankyrin repeat domain in complex with PINCH1 LIM1 domain 3hg0 
Crystal structure of a DARPin in complex with ORF49 from Lactococcal phage TP901-1 3hra 
Crystal Structure of EF0377 an Ankyrin Repeat Protein 3ixe 
Structural basis of competition between PINCH1 and PINCH2 for binding to the ankyrin repeat domain of integrin-linked kinase 3j5p 
Structure of TRPV1 ion channel determined by single particle electron cryo-microscopy 3j5q 
Structure of TRPV1 ion channel in complex with DkTx and RTX determined by single particle electron cryo-microscopy 3j5r 
Reconstruction of TRPV1 ion channel in complex with capsaicin by single particle cryo-microscopy 3j9j 
3J9J 3j9p 
3J9P 3jue 
Crystal Structure of ArfGAP and ANK repeat domain of ACAP1 3jxi 
Crystal structure of the chicken TRPV4 ankyrin repeat domain 3jxj 
Crystal structure of the chicken TRPV4 ankyrin repeat domain 3kea 
Structure function studies of vaccinia virus host-range protein K1 reveal a novel ankyrin repeat interaction surface for K1s function 3ljn 
Ankyrin repeat protein from Leishmania major 3lvq 
The crystal structure of ASAP3 in complex with Arf6 in transition state 3lvr 
The crystal structure of ASAP3 in complex with Arf6 in transition state soaked with Calcium 3nbn 
Crystal structure of a dimer of Notch Transcription Complex trimers on HES1 DNA 3noc 
Designed ankyrin repeat protein (DARPin) binders to AcrB: Plasticity of the Interface 3nog 
Designed ankyrin repeat protein (DARPin) Binders to AcrB: Plasticity of the Interface 3q9n 
In silico and in vitro co-evolution of a high affinity complementary protein-protein interface 3q9u 
In silico and in vitro co-evolution of a high affinity complementary protein-protein interface 3so8 
Crystal Structure of ANKRA 3t9k 
Crystal Structure of ACAP1 C-portion mutant S554D fused with integrin beta1 peptide 3twq 
Crystal structure of ARC4 from human Tankyrase 2 (apo form) 3twr 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human 3BP2 3tws 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human TERF1 (chimeric peptide) 3twt 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human MCL1 (chimeric peptide) 3twu 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human MCL1 3twv 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human NUMA1 (chimeric peptide) 3tww 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human LNPEP (chimeric peptide) 3twx 
Crystal structure of ARC4 from human Tankyrase 2 in complex with peptide from human FNBP1 (chimeric peptide) 3ui2 
Crystal structure of the cpSRP54 tail bound to cpSRP43 3utm 
Crystal structure of a mouse Tankyrase-Axin complex 3uxg 
Crystal structure of RFXANK 3v2o 
Crystal Structure of the Peptide Bound Complex of the Ankyrin Repeat Domains of Human ANKRA2 3v2x 
Crystal Structure of the Peptide Bound Complex of the Ankyrin Repeat Domains of Human ANKRA2 3v30 
Crystal Structure of the Peptide Bound Complex of the Ankyrin Repeat Domains of Human RFXANK 3v31 
Crystal Structure of the Peptide Bound Complex of the Ankyrin Repeat Domains of Human ANKRA2 3v79 
Structure of human Notch1 transcription complex including CSL, RAM, ANK, and MAML-1 on HES-1 promoter DNA sequence 3w9f 
Vanilloid receptor-related osmotically activated channel protein 3w9g 
Vanilloid receptor-related osmotically activated channel proteinTo be announced 3zkj 
Crystal Structure of Ankyrin Repeat and Socs Box-Containing Protein 9 (Asb9) in Complex with Elonginb and Elonginc 3zng 
Ankyrin repeat and SOCS-box protein 9 (ASB9) in complex with ElonginB and ElonginC 3zu7 
Crystal structure of a designed selected Ankyrin Repeat protein in complex with the MAP kinase ERK2 3zuv 
Crystal structure of a designed selected Ankyrin Repeat protein in complex with the phosphorylated MAP kinase ERK2 4a63 
Crystal structure of the p73-ASPP2 complex at 2.6A resolution 4atz 
Ad5 knob in complex with a designed ankyrin repeat protein 4b93 
Complex of Vamp7 cytoplasmic domain with 2nd ankyrin repeat domain of Varp 4bep 
Crystal structure of the Legionella pneumophila FIC domain-containing effector AnkX protein (apo-form) 4ber 
Crystal structure of the Legionella pneumophila FIC domain-containing effector AnkX protein in complex with cytidine monophosphate 4bes 
Crystal structure of the Legionella pneumophila FIC domain-containing effector AnkX protein in complex with cytidine monophosphate and phosphocholine 4bet 
Crystal structure of the Legionella pneumophila FIC domain-containing effector AnkX protein (inactive H229A mutant) in complex with cytidine-diphosphate-choline 4bsz 
Crystal Structure of the Yeast Ribosomal Protein Rps3 in Complex with its Chaperone Yar1 4c48 
Crystal structure of AcrB-AcrZ complex 4cym 
4CYM 4cz2 
4CZ2 4drx 
GTP-Tubulin in complex with a DARPIN 4dui 
DARPIN D1 binding to tubulin beta chain (not in complex) 4dx1 
Crystal structure of the human TRPV4 ankyrin repeat domain 4dx2 
Crystal structure of the human TRPV4 ankyrin repeat domain 4dx5 
Transport of drugs by the multidrug transporter AcrB involves an access and a deep binding pocket that are separated by a switch-loop 4dx6 
Transport of drugs by the multidrug transporter AcrB involves an access and a deep binding pocket that are separated by a switch-loop 4dx7 
Transport of drugs by the multidrug transporter AcrB involves an access and a deep binding pocket that are separated by a switch-loop 4f1p 
Crystal Structure of mutant S554D for ArfGAP and ANK repeat domain of ACAP1 4f6r 
Tubulin:Stathmin-like domain complex 4g8k 
Intact sensor domain of human RNase L in the inactive signaling state 4g8l 
Active state of intact sensor domain of human RNase L with 2-5A bound 4gmr 
Crystal Structure of Engineered Protein. Northeast Structural Genomics Consortium Target OR266. 4gpm 
Crystal Structure of Engineered Protein. Northeast Structural Genomics Consortium Target OR264. 4grg 
Crystal structure of IgE complexed with E2_79, an anti-IgE inhibitor 4hb5 
Crystal Structure of Engineered Protein. Northeast Structural Genomics Consortium Target OR267. 4hbd 
Crystal structure of KANK2 ankyrin repeats 4hi8 
Structure of integrin-linked kinase ankyrin repeat domain in complex with PINCH1 LIM1 domain collected at high energy, wavelength 0.32800 4hi9 
1.2 structure of integrin-linked kinase ankyrin repeat domain in complex with PINCH1 LIM1 domain collected at wavelength 0.91974 4hll 
Crystal structure of Artificial ankyrin repeat protein_Ank(GAG)1D4 4hna 
Kinesin motor domain in the ADP-MG-ALFX state in complex with tubulin and a DARPIN 4hqd 
Crystal Structure of Engineered Protein. Northeast Structural Genomics Consortium Target OR265. 4hrl 
Structural Basis for Eliciting a Cytotoxic Effect in HER2-Overexpressing Cancer Cells via Binding to the Extracellular Domain of HER2 4hrm 
Structural Basis for Eliciting a Cytotoxic Effect in HER2-Overexpressing Cancer Cells via Binding to the Extracellular Domain of HER2 4hrn 
Structural Basis for Eliciting a Cytotoxic Effect in HER2-Overexpressing Cancer Cells via Binding to the Extracellular Domain of HER2 4j7w 
E3_5 DARPin L86A mutant 4j8y 
E3_5 DARPin D77S mutant 4jb8 
Caspase-7 in Complex with DARPin C7_16 4k5a 
Co-crystallization with conformation-specific designed ankyrin repeat proteins explains the conformational flexibility of BCL-W 4k5b 
Co-crystallization with conformation-specific designed ankyrin repeat proteins explains the conformational flexibility of BCL-W 4k5c 
From DARPins to LoopDARPins: Novel LoopDARPin Design Allows the Selection of Low Picomolar Binders in a Single Round of Ribosome Display 4lg6 
Crystal structure of ANKRA2-CCDC8 complex 4lnu 
4LNU 4lsz 
4LSZ 4n5q 
Crystal structure of the N-terminal ankyrin repeat domain of TRPV3 4nik 
4NIK 4o1o 
Crystal Structure of RNase L in complex with 2-5A 4o1p 
Crystal Structure of RNase L in complex with 2-5A and AMP-PNP 4o60 
4O60 4oau 
Complete human RNase L in complex with biological activators. 4oav 
Complete human RNase L in complex with 2-5A (5'-ppp heptamer), AMPPCP and RNA substrate. 4ot9 
4OT9 4qfv 
4QFV 4qqi 
4QQI 4qqm 
4QQM 4rlv 
4RLV 4rly 
4RLY 4tum 
4TUM 4u8v 
4U8V 4u8y 
4U8Y 4u95 
4U95 4u96 
4U96 4uuc 
4UUC 4xcz 
4XCZ 4xd0 
4XD0 4xd1 
4XD1 4ydw 
4YDW 4ydy 
4YDY 4z68 
4Z68 4zfh 
4ZFH 4zhb 
4ZHB 5aao 
5AAO 5aar 
5AAR 5an8 
5AN8 5aq7 
5AQ7 5aq8 
5AQ8 5aq9 
5AQ9 5aqa 
5AQA 5aqb 
5AQB 5bxo 
5BXO 5bxu 
5BXU 5cbn 
5CBN 5cbo 
5CBO 5cea 
5CEA 5cec 
5CEC 5ced 
5CED 5cer 
5CER 5czy 
5CZY 5d66 
5D66 5d68 
5D68 5dnc 
5DNC 5dnd 
5DND 5dne 
5DNE 5eib 
5EIB 5eid 
5EID 5eil 
5EIL 5en5 
5EN5 5eno 
5ENO 5enp 
5ENP 5enq 
5ENQ 5enr 
5ENR 5ens 
5ENS 5ent 
5ENT 5et0 
5ET0 5et1 
5ET1 5eyl 
5EYL 5eyp 
5EYP 5fin 
5FIN 5fio 
5FIO 5hi9 
5HI9 5hkp 
5HKP 5irx 
5IRX 5irz 
5IRZ 5is0 
5IS0 5itz 
5ITZ 5iwk 
5IWK 5iwp 
5IWP 5iwr 
5IWR 5iwt 
5IWT 5ja4 
5JA4 5jhq 
5JHQ - Links (links to other resources describing this domain)
-
PFAM ank INTERPRO IPR002110

