EGF_likeEGF domain, unclasssified subfamily |
|---|
| SMART accession number: | SM00001 |
|---|---|
| Description: | - |
| Family alignment: |
There are 116150 EGF_like domains in 57116 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing EGF_like domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with EGF_like domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing EGF_like domain in the selected taxonomic class.
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
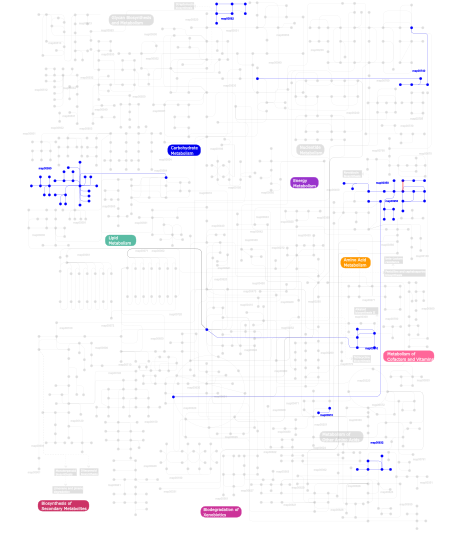
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 19.93 map04512 ECM-receptor interaction 18.82 map04510 Focal adhesion 7.75 map05222 Small cell lung cancer 7.20 map04810 Regulation of actin cytoskeleton 5.17 map04330 Notch signaling pathway 4.98 map04514 Cell adhesion molecules (CAMs) 4.43 map04360 Axon guidance 2.21 map05060 Prion disease 2.03 map04670 Leukocyte transendothelial migration 1.66 map04320 Dorso-ventral axis formation 1.48 map04060 Cytokine-cytokine receptor interaction 1.48 map04012 ErbB signaling pathway 1.29 map05212 Pancreatic cancer 1.29 map04610 Complement and coagulation cascades 1.29 map04010 MAPK signaling pathway 1.29 map05215 Prostate cancer 1.29 map04340 Hedgehog signaling pathway 1.11 map05219 Bladder cancer 1.11 map05213 Endometrial cancer 1.11 map05010 Alzheimer's disease 1.11 map05214 Glioma 1.11 map05223 Non-small cell lung cancer 1.11 map04540 Gap junction 1.11 map05218 Melanoma 0.74  map00590
map00590Arachidonic acid metabolism 0.74 map04640 Hematopoietic cell lineage 0.55  map00350
map00350Tyrosine metabolism 0.55 map04310 Wnt signaling pathway 0.55  map00740
map00740Riboflavin metabolism 0.55 map04916 Melanogenesis 0.55  map00310
map00310Lysine degradation 0.55  map00950
map00950Alkaloid biosynthesis I 0.55 map04650 Natural killer cell mediated cytotoxicity 0.37 map04350 TGF-beta signaling pathway 0.37 map04620 Toll-like receptor signaling pathway 0.18 map04520 Adherens junction 0.18 map05120 Epithelial cell signaling in Helicobacter pylori infection 0.18  map00910
map00910Nitrogen metabolism 0.18 map04662 B cell receptor signaling pathway 0.18 map05220 Chronic myeloid leukemia 0.18 map04920 Adipocytokine signaling pathway 0.18  map00632
map00632Benzoate degradation via CoA ligation 0.18  map00562
map00562Inositol phosphate metabolism 0.18 map05050 Dentatorubropallidoluysian atrophy (DRPLA) 0.18 map04210 Apoptosis 0.18 map04910 Insulin signaling pathway 0.18 map04660 T cell receptor signaling pathway 0.18 map05221 Acute myeloid leukemia 0.18 map04930 Type II diabetes mellitus This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with EGF_like domain which could be assigned to a KEGG orthologous group, and not all proteins containing EGF_like domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of EGF_like domains in PDB
PDB code Main view Title 1aut 
HUMAN ACTIVATED PROTEIN C 1c5m 
STRUCTURAL BASIS FOR SELECTIVITY OF A SMALL MOLECULE, S1-BINDING, SUB-MICROMOLAR INHIBITOR OF UROKINASE TYPE PLASMINOGEN ACTIVATOR 1dan 
COMPLEX OF ACTIVE SITE INHIBITED HUMAN BLOOD COAGULATION FACTOR VIIA WITH HUMAN RECOMBINANT SOLUBLE TISSUE FACTOR 1dva 
Crystal Structure of the Complex Between the Peptide Exosite Inhibitor E-76 and Coagulation Factor VIIA 1dx5 
Crystal structure of the thrombin-thrombomodulin complex 1ezq 
CRYSTAL STRUCTURE OF HUMAN COAGULATION FACTOR XA COMPLEXED WITH RPR128515 1f0r 
CRYSTAL STRUCTURE OF HUMAN COAGULATION FACTOR XA COMPLEXED WITH RPR208815 1f0s 
Crystal Structure of Human Coagulation Factor XA Complexed with RPR208707 1fak 
HUMAN TISSUE FACTOR COMPLEXED WITH COAGULATION FACTOR VIIA INHIBITED WITH A BPTI-MUTANT 1fax 
COAGULATION FACTOR XA INHIBITOR COMPLEX 1g2l 
FACTOR XA INHIBITOR COMPLEX 1g2m 
FACTOR XA INHIBITOR COMPLEX 1hj7 
NMR study of a pair of LDL receptor Ca2+ binding epidermal growth factor-like domains, 20 structures 1hz8 
SOLUTION STRUCTURE AND BACKBONE DYNAMICS OF A CONCATEMER OF EGF-HOMOLOGY MODULES OF THE HUMAN LOW DENSITY LIPOPROTEIN RECEPTOR 1i0u 
SOLUTION STRUCTURE AND BACKBONE DYNAMICS OF A CONCATEMER OF EGF-HOMOLOGY MODULES OF THE HUMAN LOW DENSITY LIPOPROTEIN RECEPTOR 1ioe 
Human coagulation factor Xa in complex with M55532 1iqe 
Human coagulation factor Xa in complex with M55590 1iqf 
Human coagulation factor Xa in complex with M55165 1iqg 
Human coagulation factor Xa in complex with M55159 1iqh 
Human coagulation factor Xa in complex with M55143 1iqi 
Human coagulation factor Xa in complex with M55125 1iqj 
Human coagulation factor Xa in complex with M55124 1iqk 
Human coagulation factor Xa in complex with M55113 1iql 
Human coagulation factor Xa in complex with M54476 1iqm 
Human coagulation factor Xa in complex with M54471 1iqn 
Human coagulation factor Xa in complex with M55192 1ksn 
Crystal Structure of Human Coagulation Factor XA Complexed with FXV673 1lpg 
CRYSTAL STRUCTURE OF FXA IN COMPLEX WITH 79. 1lpk 
CRYSTAL STRUCTURE OF FXA IN COMPLEX WITH 125. 1lpz 
CRYSTAL STRUCTURE OF FXA IN COMPLEX WITH 41. 1lqd 
CRYSTAL STRUCTURE OF FXA IN COMPLEX WITH 45. 1n7d 
Extracellular domain of the LDL receptor 1nfu 
CRYSTAL STRUCTURE OF HUMAN COAGULATION FACTOR XA COMPLEXED WITH RPR132747 1nfw 
CRYSTAL STRUCTURE OF HUMAN COAGULATION FACTOR XA COMPLEXED WITH RPR209685 1nfx 
CRYSTAL STRUCTURE OF HUMAN COAGULATION FACTOR XA COMPLEXED WITH RPR208944 1nfy 
CRYSTAL STRUCTURE OF HUMAN COAGULATION FACTOR XA COMPLEXED WITH RPR200095 1o5d 
Dissecting and Designing Inhibitor Selectivity Determinants at the S1 site Using an Artificial Ala190 Protease (Ala190 uPA) 1p0s 
Crystal Structure of Blood Coagulation Factor Xa in Complex with Ecotin M84R 1pfx 
PORCINE FACTOR IXA 1w0y 
tf7a_3771 complex 1w2k 
tf7a_4380 complex 1wqv 
Human Factor Viia-Tissue Factor Complexed with propylsulfonamide-D-Thr-Met-p-aminobenzamidine 1wss 
Human Factor Viia-Tissue Factor in Complex with peprid mimetic inhibitor that has two charge groups in P2 and P4 1wtg 
Human Factor Viia-Tissue Factor Complexed with ethylsulfonamide-D-biphenylalanine-Gln-p-aminobenzamidine 1wu1 
Factor Xa in complex with the inhibitor 4-[(5-chloroindol-2-yl)sulfonyl]-2-(2-methylpropyl)-1-[[5-(pyridin-4-yl) pyrimidin-2-yl]carbonyl]piperazine 1wun 
Human Factor Viia-Tissue Factor Complexed with ethylsulfonamide-D-Trp-Gln-p-aminobenzamidine 1wv7 
Human Factor Viia-Tissue Factor Complexed with ethylsulfonamide-D-5-propoxy-Trp-Gln-p-aminobenzamidine 1x7a 
Porcine Factor IXa Complexed to 1-{3-[amino(imino)methyl]phenyl}-N-[4-(1H-benzimidazol-1-yl)-2-fluorophenyl]-3-(trifluoromethyl)-1H-pyrazole-5-carboxamide 1xka 
FACTOR XA COMPLEXED WITH A SYNTHETIC INHIBITOR FX-2212A,(2S)-(3'-AMIDINO-3-BIPHENYLYL)-5-(4-PYRIDYLAMINO)PENTANOIC ACID 1xkb 
FACTOR XA COMPLEXED WITH A SYNTHETIC INHIBITOR FX-2212A,(2S)-(3'-AMIDINO-3-BIPHENYLYL)-5-(4-PYRIDYLAMINO)PENTANOIC ACID 1z6j 
Crystal Structure of a ternary complex of Factor VIIa/Tissue Factor/Pyrazinone Inhibitor 2a2q 
Complex of Active-site Inhibited Human Coagulation Factor VIIa with Human Soluble Tissue Factor in the Presence of Ca2+, Mg2+, Na+, and Zn2+ 2aei 
Crystal structure of a ternary complex of factor VIIa/tissue factor and 2-[[6-[3-(aminoiminomethyl)phenoxy]-3,5-difluro-4-[(1-methyl-3-phenylpropyl)amino]-2-pyridinyl]oxy]-benzoic acid 2aer 
Crystal Structure of Benzamidine-Factor VIIa/Soluble Tissue Factor complex. 2b7d 
Factor VIIa Inhibitors: Chemical Optimization, Preclinical Pharmacokinetics, Pharmacodynamics, and Efficacy in a Baboon Thrombosis Model 2b8o 
Crystal Structure of Glu-Gly-Arg-Chloromethyl Ketone-Factor VIIa/Soluble Tissue Factor Complex 2bo2 
EGF Domains 1,2,5 of human EMR2, a 7-TM immune system molecule, in complex with calcium. 2boh 
Crystal structure of factor Xa in complex with compound ""1"" 2bou 
EGF Domains 1,2,5 of human EMR2, a 7-TM immune system molecule, in complex with barium. 2box 
EGF Domains 1,2,5 of human EMR2, a 7-TM immune system molecule, in complex with strontium. 2c4f 
crystal structure of factor VII.stf complexed with pd0297121 2cji 
Crystal structure of a Human Factor Xa inhibitor complex 2e26 
Crystal structure of two repeat fragment of reelin 2ec9 
Crystal structure analysis of human Factor VIIa , Souluble tissue factor complexed with BCX-3607 2f9b 
Discovery of Novel Heterocyclic Factor VIIa Inhibitors 2fir 
Crystal structure of DFPR-VIIa/sTF 2flb 
Discovery of a Novel Hydroxy Pyrazole Based Factor IXa Inhibitor 2flr 
Novel 5-Azaindole Factor VIIa Inhibitors 2gy5 
Tie2 Ligand-Binding Domain Crystal Structure 2gy7 
Angiopoietin-2/Tie2 Complex Crystal Structure 2h9e 
Crystal Structure of FXa/selectide/NAPC2 ternary complex 2j2u 
CRYSTAL STRUCTURE OF A HUMAN FACTOR XA INHIBITOR COMPLEX 2j34 
CRYSTAL STRUCTURE OF A HUMAN FACTOR XA INHIBITOR COMPLEX 2j38 
CRYSTAL STRUCTURE OF A HUMAN FACTOR XA INHIBITOR COMPLEX 2j4i 
CRYSTAL STRUCTURE OF A HUMAN FACTOR XA INHIBITOR COMPLEX 2j94 
CRYSTAL STRUCTURE OF A HUMAN FACTOR XA INHIBITOR COMPLEX 2j95 
CRYSTAL STRUCTURE OF A HUMAN FACTOR XA INHIBITOR COMPLEX 2kl7 
Solution NMR Structure of the EGF-like 1 Domain of Human Fibulin-4. Northeast Structural Genomics Target HR6275 2uwl 
Selective and Dual Action Orally Active Inhibitors of Thrombin and Factor Xa 2uwo 
Selective and Dual Action Orally Active Inhibitors of Thrombin and Factor Xa 2uwp 
Factor Xa inhibitor complex 2vh0 
Structure and property based design of factor Xa inhibitors:biaryl pyrrolidin-2-ones incorporating basic heterocyclic motifs 2vh6 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with biaryl P4 motifs 2vj2 
Human Jagged-1, domains DSL and EGFs1-3 2w2m 
WT PCSK9-DELTAC BOUND TO WT EGF-A OF LDLR 2w2n 
WT PCSK9-deltaC bound to EGF-A H306Y mutant of LDLR 2w2o 
PCSK9-deltaC D374Y mutant bound to WT EGF-A of LDLR 2w2p 
PCSK9-deltaC D374A mutant bound to WT EGF-A of LDLR 2w2q 
PCSK9-deltaC D374H mutant bound to WT EGF-A of LDLR 2wyg 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with monoaryl P4 motifs 2wyj 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with monoaryl P4 motifs 2x10 
Crystal structure of the complete EphA2 ectodomain 2x11 
Crystal structure of the complete EphA2 ectodomain in complex with ephrin A5 receptor binding domain 2y7x 
The discovery of potent and long-acting oral factor Xa inhibitors with tetrahydroisoquinoline and benzazepine P4 motifs 2y7z 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with aminoindane and phenylpyrrolidine P4 motifs 2y80 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with aminoindane and phenylpyrrolidine P4 motifs 2y81 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with aminoindane and phenylpyrrolidine P4 motifs 2y82 
Structure and property based design of factor Xa inhibitors: pyrrolidin-2-ones with aminoindane and phenylpyrrolidine P4 motifs 2zp0 
Human factor viia-tissue factor complexed with benzylsulfonamide-D-ile-gln-P-aminobenzamidine 2zwl 
Human factor viia-tissue factor complexed with highly selective peptide inhibitor 2zzu 
Human Factor VIIA-Tissue Factor Complexed with ethylsulfonamide-D-5-(3-carboxybenzyloxy)-Trp-Gln-p-aminobenzamidine 3a7q 
Structural basis for specific recognition of reelin by its receptors 3bps 
PCSK9:EGF-A complex 3ela 
Crystal structure of active site inhibited coagulation factor VIIA mutant in complex with soluble tissue factor 3ens 
Crystal structure of human FXA in complex with methyl (2Z)-3-[(3-chloro-1H-indol-7-yl)amino]-2-cyano-3-{[(3S)-2-oxo-1-(2-oxo-2-pyrrolidin-1-ylethyl)azepan-3-yl]amino}acrylate 3fl7 
Crystal structure of the human ephrin A2 ectodomain 3gcw 
PCSK9:EGFA(H306Y) 3gcx 
PCSK9:EGFA (pH 7.4) 3hpt 
Crystal structure of human FxA in complex with (S)-2-cyano-1-(2-methylbenzofuran-5-yl)-3-(2-oxo-1-(2-oxo-2-(pyrrolidin-1-yl)ethyl)azepan-3-yl)guanidine 3k9x 
X-ray crystal structure of human fxa in complex with (S)-N-((2-METHYLBENZOFURAN-5-YLAMINO)(2-OXO-1-(2-OXO-2- (PYRROLIDIN-1-YL)ETHYL)AZEPAN-3- YLAMINO)METHYLENE)NICOTINAMIDE 3m0c 
The X-ray Crystal Structure of PCSK9 in Complex with the LDL receptor 3mbw 
Crystal structure of the human ephrin A2 LBD and CRD domains in complex with ephrin A1 3mx0 
Crystal Structure of EphA2 ectodomain in complex with ephrin-A5 3ooi 
Crystal Structure of Human Histone-Lysine N-methyltransferase NSD1 SET domain in Complex with S-adenosyl-L-methionine 3p5b 
The structure of the LDLR/PCSK9 complex reveals the receptor in an extended conformation 3p5c 
The structure of the LDLR/PCSK9 complex reveals the receptor in an extended conformation 3s8v 
Crystal structure of LRP6-Dkk1 complex 3s8z 
Crystal structure of LRP6-E3E4 3sw2 
X-ray crystal structure of human FXA in complex with 6-chloro-N-((3S)-2-oxo-1-(2-oxo-2-((5S)-8-oxo-5,6-dihydro-1H-1,5-methanopyrido[1,2-a][1,5]diazocin-3(2H,4H,8H)-yl)ethyl)piperidin-3-yl)naphthalene-2-sulfonamide 3th2 
Mg2+ Is Required for Optimal Folding of the Gamma-Carboxyglutamic Acid (Gla) Domains of Vitamin K-Dependent Clotting Factors At Physiological Ca2+ 3th3 
Mg2+ Is Required for Optimal Folding of the Gamma-Carboxyglutamic Acid (Gla) Domains of Vitamin K-Dependent Clotting Factors At Physiological Ca2+ 3th4 
Mg2+ Is Required for Optimal Folding of the Gamma-Carboxyglutamic Acid (Gla) Domains of Vitamin K-Dependent Clotting Factors At Physiological Ca2+ 3v65 
Crystal structure of agrin and LRP4 complex 3zyj 
NetrinG1 in complex with NGL1 4a7i 
Factor Xa in complex with a potent 2-amino-ethane sulfonamide inhibitor 4aqs 
Laminin beta1 LN-LE1-4 structure 4bti 
factor Xa in complex with the dual thrombin-FXa inhibitor 58. 4btt 
factor Xa in complex with the dual thrombin-FXa inhibitor 31. 4btu 
Factor Xa in complex with the dual thrombin-FXa inhibitor 57. 4bxs 
Crystal Structure of the Prothrombinase Complex from the Venom of Pseudonaja Textilis 4bxw 
Crystal Structure of the Prothrombinase Complex from the Venom of Pseudonaja Textilis 4cbz 
Notch ligand, Jagged-1, contains an N-terminal C2 domain 4cc0 
Notch ligand, Jagged-1, contains an N-terminal C2 domain 4cc1 
Notch ligand, Jagged-1, contains an N-terminal C2 domain 4d90 
Crystal Structure of Del-1 EGF domains 4ibl 
Rubidium Sites in Blood Coagulation Factor VIIa 4k0v 
Structural basis for angiopoietin-1 mediated signaling initiation 4um8 
4UM8 4um9 
4UM9 4xbm 
4XBM 4y6d 
4Y6D 4y71 
4Y71 4y76 
4Y76 4y79 
4Y79 4y7a 
4Y7A 4y7b 
4Y7B 4ylq 
4YLQ 4yz8 
4YZ8 4zh8 
4ZH8 4zha 
4ZHA 4zma 
4ZMA 5bo1 
5BO1 5fm9 
5FM9 5fma 
5FMA 5jb8 
5JB8 5jb9 
5JB9 5jba 
5JBA 5jbb 
5JBB 5jbc 
5JBC 5lsu 
5LSU

