The domain within your query sequence starts at position 116 and ends at position 821; the E-value for the MYSc domain shown below is < 1e-12.
MNPPKFSKVEDMAELTCLNEASVLHNLKDRYYSGLIYTYSGLFCVVINPYKNLPIYSENI IEMYRGKKRHEMPPHIYAISESAYRCMLQDREDQSILCTGESGAGKTENTKKVIQYLAHV ASSHKGRKDHNIPGELERQLLQANPILESFGNAKTVKNDNSSRFGKFIRINFDVTGYIVG ANIETYLLEKSRAVRQAKDERTFHIFYQLLSGAGEHLKSDLLLEGFNNYRFLSNGYIPIP GQQDKDNFQETMEAMHIMGFSHEEILSMLKVVSSVLQFGNISFKKERNTDQASMPENTVA QKLCHLLGMNVMEFTRAILTPRIKVGRDYVQKAQTKEQADFAVEALAKATYERLFRWLVH RINKALDRTKRQGASFIGILDIAGFEIFELNSFEQLCINYTNEKLQQLFNHTMFILEQEE YQREGIEWNFIDFGLDLQPCIDLIERPANPPGVLALLDEECWFPKATDKTFVEKLVQEQG SHSKFQKPRQLKDKADFCIIHYAGKVDYKADEWLMKNMDPLNDNVATLLHQSSDRFVAEL WKDVDRIVGLDQVTGMTETAFGSAYKTKKGMFRTVGQLYKESLTKLMATLRNTNPNFVRC IIPNHEKRAGKLDPHLVLDQLRCNGVLEGIRICRQGFPNRIVFQEFRQRYEILTPNAIPK GFMDGKQACERMIRALELDPNLYRIGQSKIFFRAGVLAHLEEERDL
MYScMyosin. Large ATPases. |
|---|
| SMART accession number: | SM00242 |
|---|---|
| Description: | ATPase; molecular motor. Muscle contraction consists of a cyclical interaction between myosin and actin. The core of the myosin structure is similar in fold to that of kinesin. |
| Interpro abstract (IPR001609): | Muscle contraction is caused by sliding between the thick and thin filaments of the myofibril. Myosin is a major component of thick filaments and exists as a hexamer of 2 heavy chains [ (PUBMED:1939027) ], 2 alkali light chains, and 2 regulatory light chains. The heavy chain can be subdivided into the N-terminal globular head and the C-terminal coiled-coil rod-like tail, although some forms have a globular region in their C-terminal. There are many cell-specific isoforms of myosin heavy chains, coded for by a multi-gene family [ (PUBMED:2806546) ]. Myosin interacts with actin to convert chemical energy, in the form of ATP, to mechanical energy [ (PUBMED:3540939) ]. The 3-D structure of the head portion of myosin has been determined [ (PUBMED:8316857) ] and a model for actin-myosin complex has been constructed [ (PUBMED:8316858) ]. The globular head is well conserved, some highly-conserved regions possibly relating to functional and structural domains [ (PUBMED:6576334) ]. The rod-like tail starts with an invariant proline residue, and contains many repeats of a 28 residue region, interrupted at 4 regularly-spaced points known as skip residues. Although the sequence of the tail is not well conserved, the chemical character is, hydrophobic, charged and skip residues occuring in a highly ordered and repeated fashion [ (PUBMED:6576334) ]. |
| GO component: | myosin complex (GO:0016459) |
| GO function: | ATP binding (GO:0005524), motor activity (GO:0003774) |
| Family alignment: |
There are 33491 MYSc domains in 33449 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing MYSc domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with MYSc domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing MYSc domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Binding / catalysis: ATP-hydrolysis, actin-binding
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Hasson T, Mooseker MS
- The growing family of myosin motors and their role in neurons and sensory cells.
- Curr Opin Neurobiol. 1997; 7: 615-23
- Display abstract
Biochemical and physiological evidence has suggested that myosins, both conventional and unconventional, are critical for neurosensory activities. In the past few years, this premise has been supported by genetic evidence that has shown that unconventional myosins are essential for the proper functioning of neurons, retina and the sensory cells of the inner ear.
- Cope MJ, Whisstock J, Rayment I, Kendrick-Jones J
- Conservation within the myosin motor domain: implications for structure and function.
- Structure. 1996; 4: 969-87
- Display abstract
BACKGROUND: Myosins are motors that use energy supplied by ATP to travel along actin filaments. The structure of myosin is known, but the actin-binding site is not well defined, and the mechanisms by which actin activates ATP hydrolysis by myosin, and myosin moves relative to the actin filament, developing force, are not fully understood. Previous phylogenetic analyses of the motor domain of myosins have identified up to twelve classes. We set out to analyse the positions of conserved residues within this domain in detail, and relate the conserved residues to the myosin structure. RESULTS: Our analysis indicates that there are at least thirteen myosin classes. Conserved residues in the motor domain have been positioned within the framework provided by the recent crystal structures, thus helping to define those residues involved in actin and ATP binding, in hydrolysis and in conformational change. This has revealed remarkably poor overall conservation at the site thought to be involved in actin binding, but several highly conserved residues have been identified that may be functionally important. CONCLUSIONS: Information from such a sequence analysis is a useful tool in the further interpretation of X-ray structures. It allows the position of crucial residues from other members of a superfamily to be determined within the framework provided by the known structures and the functional significance of conserved or mutated residues to be assessed.
- Smith CA, Rayment I
- X-ray structure of the magnesium(II).ADP.vanadate complex of the Dictyostelium discoideum myosin motor domain to 1.9 A resolution.
- Biochemistry. 1996; 35: 5404-17
- Display abstract
The structure of the vanadate-trapped ADP complex of a truncated head of Dictyostelium myosin II consisting of residues Asp 2-Asn 762 has been determined by molecular replacement at 1.9 A resolution and refined to a crystallographic R-factor of 19.4%. The crystals belong to the orthorhombic space group C2221 where a = 84.50 A, b = 145.4 A, and c = 152.8 A. The conformation of the protein is similar to that of MgADP.AlF4.SlDc [Fisher, A.J., et al. (1995) Biochemistry 34, 8960-8972]. The nucleotide binding site contains a complex between MgADP and vanadate where MgADP exhibits a very similar conformation to that seen in previous complexes. The vanadate ion adopts a trigonal bipyramidal coordination. The three equatorial oxygen ligands are fairly short, average 1.7 A, relative to a single bond distance of approximately 1.8 A and are coordinated to the magnesium ion, N zeta of Lys 185, and five other protein ligands. The apical coordination to the vanadate ion is filled by a terminal oxygen on the beta-phosphate of ADP and a water molecule at bond distances of 2.1 and 2.3 A, respectively. The long length of the apical bonds suggests that the bond order is considerably less than unity. This structure confirms the earlier suggestion that vanadate is a model for the transition state of ATP hydrolysis and thus provides insight into those factors that are responsible for catalysis. In particular, it shows that the protein ligands and water structure surrounding the gamma-phosphate pocket are oriented to stabilize a water molecule in an appropriate position for in-line nucleophilic attack on the gamma-phosphorus of ATP. This structure reveals also an orientation of the COOH-terminal region beyond Thr 688 which is very different from that observed in either MgADP.BeFx.SlDc or chicken skeletal myosin subfragment 1. This is consistent with the COOH-terminal region of the molecule playing an important role in the transduction of chemical energy of hydrolysis of ATP into mechanical movement.
- Fisher AJ et al.
- X-ray structures of the myosin motor domain of Dictyostelium discoideum complexed with MgADP.BeFx and MgADP.AlF4-.
- Biochemistry. 1995; 34: 8960-72
- Display abstract
The three-dimensional structures of the truncated myosin head from Dictyostelium discoideum myosin II complexed with beryllium and aluminum fluoride and magnesium ADP are reported at 2.0 and 2.6 A resolution, respectively. Crystals of the beryllium fluoride-MgADP complex belong to space group P2(1)2(1)2 with unit cell parameters of a = 105.3 A, b = 182.6 A, and c = 54.7 A, whereas the crystals of the aluminum fluoride complex belong to the orthorhombic space group C222(1) with unit cell dimensions of a = 87.9 A, b = 149.0 A, and c = 153.8 A. Chemical modification was not necessary to obtain these crystals. These structures reveal the location of the nucleotide complexes and define the amino acid residues that form the active site. The tertiary structure of the protein complexed with MgADP.BeFx is essentially identical to that observed previously in the three-dimensional model of chicken skeletal muscle myosin subfragment-1 in which no nucleotide was present. By contrast, the complex with MgADP.AlF4- exhibits significant domain movements. The structures suggest that the MgADP.BeFx complex mimics the ATP bound state and the MgADP.AlF4- complex is an analog of the transition state for hydrolysis. The domain movements observed in the MgADP.AlF4- complex indicate that myosin undergoes a conformational change during hydrolysis that is not associated with the nucleotide binding pocket but rather occurs in the COOH-terminal segment of the myosin motor domain.
- Smith CA, Rayment I
- X-ray structure of the magnesium(II)-pyrophosphate complex of the truncated head of Dictyostelium discoideum myosin to 2.7 A resolution.
- Biochemistry. 1995; 34: 8973-81
- Display abstract
The structure of the magnesium pyrophosphate complex of the truncated head of Dictyostelium myosin has been determined by molecular replacement at 2.7 A resolution and refined to a crystallographic R-factor of 16.0%. The crystals belong to the orthorhombic space group P2(1)2(1)2, where a = 105.2 A, b = 182.1 A, and c = 54.5 A. The conformation of the protein around the magnesium pyrophosphate is very similar to that seen when magnesium ADP-beryllium fluoride binds in the active site. The latter complex mimics the binding of ATP prior to hydrolysis. The pyrophosphate molecule occupies the beta- and gamma-phosphate sites, where the two phosphorus atoms are in the same positions as the beta-phosphate and the BeFx moiety of the beryllium fluoride-trapped ADP. The surrounding active site residues are almost perfectly superimposable in the two structures and the hydrogen-bonding interactions that the PPi makes with the protein are essentially identical. The similarity between the MgPPi and MgADP.BeFx complex with S1Dc suggests that the conformational change, which occurs when ATP binds to actomyosin and which reduces the affinity of myosin for actin, is caused by the binding of the gamma- and beta-phosphate groups of the nucleotide. This then implies that the role of the remainder of the substrate is to increase the binding affinity for myosin and thus to drive the equilibrium toward dissociation of myosin from actin.
- Rayment I, Holden HM
- The three-dimensional structure of a molecular motor.
- Trends Biochem Sci. 1994; 19: 129-34
- Display abstract
Myosin is one of only three proteins known to convert chemical energy into mechanical work. Although the chemical, kinetic and physiological characteristics of this protein have been studied extensively, it has been difficult to define its molecular basis of movement. With the recent X-ray structural determination of the myosin head, however, it is now possible to put forward a hypothesis on how myosin might function as a molecular motor.
- Xie X et al.
- Structure of the regulatory domain of scallop myosin at 2.8 A resolution.
- Nature. 1994; 368: 306-12
- Display abstract
The regulatory domain of scallop myosin is a three-chain protein complex that switches on this motor in response to Ca2+ binding. This domain has been crystallized and the structure solved to 2.8 A resolution. Side-chain interactions link the two light chains in tandem to adjacent segments of the heavy chain bearing the IQ-sequence motif. The Ca(2+)-binding site is a novel EF-hand motif on the essential light chain and is stabilized by linkages involving the heavy chain and both light chains, accounting for the requirement of all three chains for Ca2+ binding and regulation in the intact myosin molecule.
- Rayment I et al.
- Three-dimensional structure of myosin subfragment-1: a molecular motor.
- Science. 1993; 261: 50-8
- Display abstract
Directed movement is a characteristic of many living organisms and occurs as a result of the transformation of chemical energy into mechanical energy. Myosin is one of three families of molecular motors that are responsible for cellular motility. The three-dimensional structure of the head portion of myosin, or subfragment-1, which contains both the actin and nucleotide binding sites, is described. This structure of a molecular motor was determined by single crystal x-ray diffraction. The data provide a structural framework for understanding the molecular basis of motility.
- Rayment I et al.
- Structure of the actin-myosin complex and its implications for muscle contraction.
- Science. 1993; 261: 58-65
- Display abstract
Muscle contraction consists of a cyclical interaction between myosin and actin driven by the concomitant hydrolysis of adenosine triphosphate (ATP). A model for the rigor complex of F actin and the myosin head was obtained by combining the molecular structures of the individual proteins with the low-resolution electron density maps of the complex derived by cryo-electron microscopy and image analysis. The spatial relation between the ATP binding pocket on myosin and the major contact area on actin suggests a working hypothesis for the crossbridge cycle that is consistent with previous independent structural and biochemical studies.
- Disease (disease genes where sequence variants are found in this domain)
-
SwissProt sequences and OMIM curated human diseases associated with missense mutations within the MYSc domain.
Protein Disease Myosin-9 (P35579) (SMART) OMIM:155100: May-Hegglin anomaly
OMIM:160775: May-Hegglin anomaly
OMIM:155100: Fechtner syndrome
OMIM:153640: Sebastian syndrome
OMIM:605249: Deafness, autosomal dominant 17
OMIM:603622:Unconventional myosin-VIIa (Q13402) (SMART) OMIM:276903: Usher syndrome, type 1B ; Deafness, autosomal recessive 2, neurosensory
OMIM:600060: Deafness, autosomal dominant 11, neurosensory
OMIM:601317:UNKNOWN (SMART) OMIM:160760: Cardiomyopathy, familial hypertrophic, 1
OMIM:192600: ?Central core disease, one form - Metabolism (metabolic pathways involving proteins which contain this domain)
-
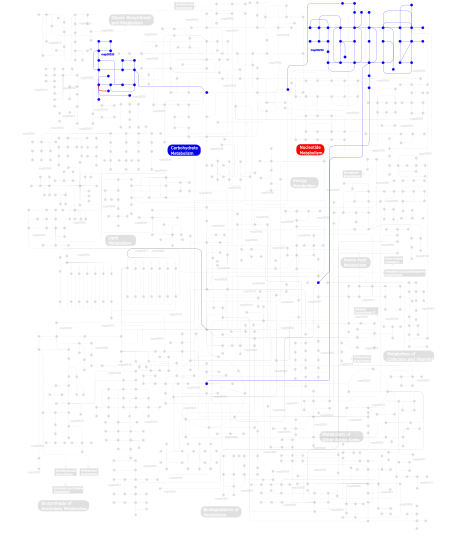
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 91.80 map04530 Tight junction 5.74  map00230
map00230Purine metabolism 2.46  map00530
map00530Aminosugars metabolism This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with MYSc domain which could be assigned to a KEGG orthologous group, and not all proteins containing MYSc domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of MYSc domains in PDB
PDB code Main view Title 1b7t 
MYOSIN DIGESTED BY PAPAIN 1br1 
SMOOTH MUSCLE MYOSIN MOTOR DOMAIN-ESSENTIAL LIGHT CHAIN COMPLEX WITH MGADP.ALF4 BOUND AT THE ACTIVE SITE 1br2 
SMOOTH MUSCLE MYOSIN MOTOR DOMAIN COMPLEXED WITH MGADP.ALF4 1br4 
SMOOTH MUSCLE MYOSIN MOTOR DOMAIN-ESSENTIAL LIGHT CHAIN COMPLEX WITH MGADP.BEF3 BOUND AT THE ACTIVE SITE 1d0x 
DICTYOSTELIUM MYOSIN S1DC (MOTOR DOMAIN FRAGMENT) COMPLEXED WITH M-NITROPHENYL AMINOETHYLDIPHOSPHATE BERYLLIUM TRIFLUORIDE. 1d0y 
DICTYOSTELIUM MYOSIN S1DC (MOTOR DOMAIN FRAGMENT) COMPLEXED WITH O-NITROPHENYL AMINOETHYLDIPHOSPHATE BERYLLIUM FLUORIDE. 1d0z 
DICTYOSTELIUM MYOSIN S1DC (MOTOR DOMAIN FRAGMENT) COMPLEXED WITH P-NITROPHENYL AMINOETHYLDIPHOSPHATE BERYLLIUM TRIFLUORIDE. 1d1a 
DICTYOSTELIUM MYOSIN S1DC (MOTOR DOMAIN FRAGMENT) COMPLEXED WITH O,P-DINITROPHENYL AMINOETHYLDIPHOSPHATE BERYLLIUM TRIFLUORIDE. 1d1b 
DICTYOSTELIUM MYOSIN S1DC (MOTOR DOMAIN FRAGMENT) COMPLEXED WITH O,P-DINITROPHENYL AMINOPROPYLDIPHOSPHATE BERYLLIUM TRIFLUORIDE. 1d1c 
DICTYOSTELIUM MYOSIN S1DC (MOTOR DOMAIN FRAGMENT) COMPLEXED WITH N-METHYL-O-NITROPHENYL AMINOETHYLDIPHOSPHATE BERYLLIUM TRIFLUORIDE. 1dfk 
NUCLEOTIDE-FREE SCALLOP MYOSIN S1-NEAR RIGOR STATE 1dfl 
SCALLOP MYOSIN S1 COMPLEXED WITH MGADP:VANADATE-TRANSITION STATE 1fmv 
CRYSTAL STRUCTURE OF THE APO MOTOR DOMAIN OF DICTYOSTELLIUM MYOSIN II 1fmw 
CRYSTAL STRUCTURE OF THE MGATP COMPLEX FOR THE MOTOR DOMAIN OF DICTYOSTELIUM MYOSIN II 1g8x 
STRUCTURE OF A GENETICALLY ENGINEERED MOLECULAR MOTOR 1i84 
CRYO-EM STRUCTURE OF THE HEAVY MEROMYOSIN SUBFRAGMENT OF CHICKEN GIZZARD SMOOTH MUSCLE MYOSIN WITH REGULATORY LIGHT CHAIN IN THE DEPHOSPHORYLATED STATE. ONLY C ALPHAS PROVIDED FOR REGULATORY LIGHT CHAIN. ONLY BACKBONE ATOMS PROVIDED FOR S2 FRAGMENT. 1jwy 
CRYSTAL STRUCTURE OF THE DYNAMIN A GTPASE DOMAIN COMPLEXED WITH GDP, DETERMINED AS MYOSIN FUSION 1jx2 
CRYSTAL STRUCTURE OF THE NUCLEOTIDE-FREE DYNAMIN A GTPASE DOMAIN, DETERMINED AS MYOSIN FUSION 1kk7 
SCALLOP MYOSIN IN THE NEAR RIGOR CONFORMATION 1kk8 
SCALLOP MYOSIN (S1-ADP-BeFx) IN THE ACTIN-DETACHED CONFORMATION 1kqm 
SCALLOP MYOSIN S1-AMPPNP IN THE ACTIN-DETACHED CONFORMATION 1kwo 
SCALLOP MYOSIN S1-ATPgammaS-p-PDM IN THE ACTIN-DETACHED CONFORMATION 1l2o 
SCALLOP MYOSIN S1-ADP-p-PDM IN THE ACTIN-DETACHED CONFORMATION 1lkx 
MOTOR DOMAIN OF MYOE, A CLASS-I MYOSIN 1lvk 
X-RAY CRYSTAL STRUCTURE OF THE MG (DOT) 2'(3')-O-(N-METHYLANTHRANILOYL) NUCLEOTIDE BOUND TO DICTYOSTELIUM DISCOIDEUM MYOSIN MOTOR DOMAIN 1m8q 
Molecular Models of Averaged Rigor Crossbridges from Tomograms of Insect Flight Muscle 1mma 
X-RAY STRUCTURES OF THE MGADP, MGATPGAMMAS, AND MGAMPPNP COMPLEXES OF THE DICTYOSTELIUM DISCOIDEUM MYOSIN MOTOR DOMAIN 1mmd 
TRUNCATED HEAD OF MYOSIN FROM DICTYOSTELIUM DISCOIDEUM COMPLEXED WITH MGADP-BEF3 1mmg 
X-RAY STRUCTURES OF THE MGADP, MGATPGAMMAS, AND MGAMPPNP COMPLEXES OF THE DICTYOSTELIUM DISCOIDEUM MYOSIN MOTOR DOMAIN 1mmn 
X-RAY STRUCTURES OF THE MGADP, MGATPGAMMAS, AND MGAMPPNP COMPLEXES OF THE DICTYOSTELIUM DISCOIDEUM MYOSIN MOTOR DOMAIN 1mnd 
TRUNCATED HEAD OF MYOSIN FROM DICTYOSTELIUM DISCOIDEUM COMPLEXED WITH MGADP-ALF4 1mne 
TRUNCATED HEAD OF MYOSIN FROM DICTYOSTELIUM DISCOIDEUM COMPLEXED WITH MG-PYROPHOSPHATE 1mvw 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o18 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o19 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1a 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1b 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1c 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1d 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1e 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1f 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1g 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1oe9 
Crystal structure of Myosin V nucleotide-free 1qvi 
Crystal structure of scallop myosin S1 in the pre-power stroke state to 2.6 Angstrom resolution: flexibility and function in the head 1s5g 
Structure of Scallop myosin S1 reveals a novel nucleotide conformation 1sr6 
Structure of nucleotide-free scallop myosin S1 1vom 
COMPLEX BETWEEN DICTYOSTELIUM MYOSIN AND MGADP AND VANADATE AT 1.9A RESOLUTION 1w7i 
Crystal Structure Of Myosin V Motor Without nucleotide soaked in 10 mM MgADP 1w7j 
Crystal Structure Of Myosin V Motor With Essential Light Chain + ADP-BeFx - Near Rigor 1w8j 
Crystal Structure Of Myosin V Motor Domain - Nucleotide-Free 1w9i 
Myosin II Dictyostelium discoideum motor domain S456Y bound with MgADP-BeFx 1w9j 
Myosin II Dictyostelium discoideum motor domain S456Y bound with MgADP-AlF4 1w9k 
Dictyostelium discoideum Myosin II motor domain S456E with bound MgADP-BeFx 1w9l 
Myosin II Dictyostelium discoideum motor domain S456E bound with MgADP-AlF4 1yv3 
The structural basis of blebbistatin inhibition and specificity for myosin II 2aka 
Structure of the nucleotide-free myosin II motor domain from Dictyostelium discoideum fused to the GTPase domain of dynamin 1 from Rattus norvegicus 2bkh 
Myosin VI nucleotide-free (MDInsert2) crystal structure. 2bki 
Myosin VI nucleotide-free (MDinsert2-IQ) crystal structure 2dfs 
3-D structure of Myosin-V inhibited state 2ec6 
Placopecten Striated Muscle Myosin II 2jhr 
Crystal structure of myosin-2 motor domain in complex with ADP- metavanadate and pentabromopseudilin 2jj9 
Crystal structure of myosin-2 in complex with ADP-metavanadate 2mys 
MYOSIN SUBFRAGMENT-1, ALPHA CARBON COORDINATES ONLY FOR THE TWO LIGHT CHAINS 2os8 
Rigor-like structures of muscle myosins reveal key mechanical elements in the transduction pathways of this allosteric motor 2otg 
Rigor-like structures of muscle myosins reveal key mechanical elements in the transduction pathways of this allosteric motor 2v26 
Myosin VI (MD) pre-powerstroke state (Mg.ADP.VO4) 2vas 
Myosin VI (MD-insert2-CaM, Delta-Insert1) Post-rigor state 2vb6 
Myosin VI (MD-insert2-CaM, Delta Insert1) Post-rigor state (crystal form 2) 2w4a 
Isometrically contracting insect asynchronous flight muscle 2w4g 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick stretch step 2w4h 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick release step 2w4t 
Isometrically contracting insect asynchronous flight muscle 2w4v 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick release step 2w4w 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick stretch step 2x51 
M6 delta Insert1 2x9h 
CRYSTAL STRUCTURE OF MYOSIN-2 MOTOR DOMAIN IN COMPLEX WITH ADP- METAVANADATE AND PENTACHLOROCARBAZOLE 2xel 
Molecular Mechanism of Pentachloropseudilin Mediated Inhibition of Myosin Motor Activity 2xo8 
Crystal Structure of Myosin-2 in Complex with Tribromodichloropseudilin 2y0r 
Structural basis for the allosteric interference of myosin function by mutants G680A and G680V of Dictyostelium myosin-2 2y8i 
Structural basis for the allosteric interference of myosin function by mutants G680A and G680V of Dictyostelium myosin-2 2y9e 
Structural basis for the allosteric interference of myosin function by mutants G680A and G680V of Dictyostelium myosin-2 3bz7 
Crystal Structures of (S)-(-)-Blebbistatin Analogs bound to Dictyostelium discoideum myosin II 3bz8 
Crystal Structures of (S)-(-)-Blebbistatin Analogs bound to Dictyostelium discoideum myosin II 3bz9 
Crystal Structures of (S)-(-)-Blebbistatin Analogs bound to Dictyostelium discoideum myosin II 3dtp 
Tarantula heavy meromyosin obtained by flexible docking to Tarantula muscle thick filament Cryo-EM 3D-MAP 3i5f 
Crystal structure of squid MG.ADP myosin S1 3i5g 
Crystal structure of rigor-like squid myosin S1 3i5h 
The crystal structure of rigor like squid myosin S1 in the absence of nucleotide 3i5i 
The crystal structure of squid myosin S1 in the presence of SO4 2- 3j04 
EM structure of the heavy meromyosin subfragment of Chick smooth muscle Myosin with regulatory light chain in phosphorylated state 3jax 
3JAX 3jbh 
3JBH 3l9i 
Myosin VI nucleotide-free (mdinsert2) L310G mutant crystal structure 3mjx 
Crystal structure of myosin-2 motor domain in complex with ADP-Metavanadate and blebbistatin 3mkd 
Crystal structure of myosin-2 dictyostelium discoideum motor domain S456Y mutant in complex with adp-orthovanadate 3mnq 
Crystal structure of myosin-2 motor domain in complex with ADP-metavanadate and resveratrol 3myh 
Insights into the Importance of Hydrogen Bonding in the Gamma-Phosphate Binding Pocket of Myosin: Structural and Functional Studies of Ser236 3myk 
Insights into the Importance of Hydrogen Bonding in the Gamma-Phosphate Binding Pocket of Myosin: Structural and Functional Studies of Ser236 3myl 
Insights into the Importance of Hydrogen Bonding in the Gamma-Phosphate Binding Pocket of Myosin: Structural and Functional Studies of Ser236 4a7f 
Structure of the Actin-Tropomyosin-Myosin Complex (rigor ATM 3) 4a7h 
Structure of the Actin-Tropomyosin-Myosin Complex (rigor ATM 2) 4a7l 
Structure of the Actin-Tropomyosin-Myosin Complex (rigor ATM 1) 4ae3 
Crystal structure of ammosamide 272:myosin-2 motor domain complex 4anj 
MYOSIN VI (MDinsert2-GFP fusion) PRE-POWERSTROKE STATE (MG.ADP.AlF4) 4byf 
Crystal structure of human Myosin 1c in complex with calmodulin in the pre-power stroke state 4db1 
Cardiac human myosin S1dC, beta isoform complexed with Mn-AMPPNP 4dbp 
Myosin VI nucleotide-free (MDINSERT2) D179Y crystal structure 4dbq 
MYOSIN VI D179Y (MD-INSERT2-CAM, DELTA-INSERT1) post-rigor state 4dbr 
Myosin VI D179Y (MD) pre-powerstroke state 4e7s 
Myosin VI D23R I24R R569E (MD) pre-powerstroke state 4e7z 
Myosin VI (MD) pre-powerstroke state, P21 crystal form 4l79 
Crystal Structure of nucleotide-free Myosin 1b residues 1-728 with bound Calmodulin 4p7h 
Structure of Human beta-Cardiac Myosin Motor Domain::GFP chimera 4pa0 
4PA0 4pd3 
4PD3 4pfo 
4PFO 4pfp 
4PFP 4pjj 
4PJJ 4pjk 
4PJK 4pjl 
4PJL 4pjm 
4PJM 4pjn 
4PJN 4pk4 
4PK4 4qbd 
4QBD 4zg4 
4ZG4 4zlk 
4ZLK 5hmp 
5HMP 5i0h 
5I0H 5i0i 
5I0I 5jlh 
5JLH 5kg8 
5KG8 5m05 
5M05 5t45 
5T45 - Links (links to other resources describing this domain)
-
PFAM myosin_head INTERPRO IPR001609

