PP2CcSerine/threonine phosphatases, family 2C, catalytic domain |
|---|
| SMART accession number: | SM00332 |
|---|---|
| Description: | The protein architecture and deduced catalytic mechanism of PP2C phosphatases are similar to the PP1, PP2A, PP2B family of protein Ser/Thr phosphatases, with which PP2C shares no sequence similarity. |
| Family alignment: |
There are 66722 PP2Cc domains in 66644 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing PP2Cc domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with PP2Cc domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing PP2Cc domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Cellular role: signalling
Binding / catalysis: protein phosphatase, serine-specific phosphatase, threonine-specific phosphatase - Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Adler E, Donella-Deana A, Arigoni F, Pinna LA, Stragler P
- Structural relationship between a bacterial developmental protein and eukaryotic PP2C protein phosphatases.
- Mol Microbiol. 1997; 23: 57-62
- Display abstract
Bacillus subtilis SpoIIE is a Ser protein phosphatase whose action on the phosphoprotein SpoIIAA triggers the cell type-specific activation of a sporulation transcription factor. Here we report that SpoIIE displays sequence similarity to the PP2C family of eukaryotic Ser/Thr protein phosphatases, and that residues common to these proteins are required for the function of both SpoIIE and TPD1, a yeast PP2C. These findings suggest that SpoIIE and the PP2C protein phosphatases are structurally related, and reveal a striking formal similarity between the SpoIIAA regulatory circuit and that of mammalian mitochondrial pyruvate dehydrogenase. This similarity may reflect an evolutionarily conserved mechanism of biological regulation based on the interplay of His protein kinase-like Ser kinases and PP2C-like protein phosphatases.
- Cohen PT
- Novel protein serine/threonine phosphatases: variety is the spice of life.
- Trends Biochem Sci. 1997; 22: 245-51
- Display abstract
The dephosphorylation of proteins on serine and threonine residues is a major mechanism of cellular regulation. Many novel protein serine/threonine phosphatases in the PPP family have recently been discovered and the insights that have been gained into their different functions are summarised in this review.
- Bork P, Brown NP, Hegyi H, Schultz J
- The protein phosphatase 2C (PP2C) superfamily: detection of bacterial homologues.
- Protein Sci. 1996; 5: 1421-5
- Display abstract
A thorough sequence analysis of the various members of the eukaryotic protein serine/threonine phosphatase 2C (PP2C) family revealed the conservation of 11 motifs. These motifs could be identified in numerous other sequences, including fungal adenylate cyclases that are predicted to contain a functionally active PP2C domain, and a family of prokaryotic serine/threonine phosphatases including SpoIIE. Phylogenetic analysis of all the proteins indicates a widespread sequence family for which a considerable number of isoenzymes can be inferred.
- Das AK, Helps NR, Cohen PT, Barford D
- Crystal structure of the protein serine/threonine phosphatase 2C at 2.0 A resolution.
- EMBO J. 1996; 15: 6798-809
- Display abstract
Protein phosphatase 2C (PP2C) is a Mn2+- or Mg2+-dependent protein Ser/Thr phosphatase that is essential for regulating cellular stress responses in eukaryotes. The crystal structure of human PP2C reveals a novel protein fold with a catalytic domain composed of a central beta-sandwich that binds two manganese ions, which is surrounded by alpha-helices. Mn2+-bound water molecules at the binuclear metal centre coordinate the phosphate group of the substrate and provide a nucleophile and general acid in the dephosphorylation reaction. Our model presents a framework for understanding not only the classical Mn2+/Mg2+-dependent protein phosphatases but also the sequence-related domains of mitochondrial pyruvate dehydrogenase phosphatase, the Bacillus subtilus phosphatase SpoIIE and a 300-residue domain within yeast adenyl cyclase. The protein architecture and deduced catalytic mechanism are strikingly similar to the PP1, PP2A, PP2B family of protein Ser/Thr phosphatases, with which PP2C shares no sequence similarity, suggestive of convergent evolution of protein Ser/Thr phosphatases.
- Mumby MC, Walter G
- Protein serine/threonine phosphatases: structure, regulation, and functions in cell growth.
- Physiol Rev. 1993; 73: 673-99
- Display abstract
It is clear that much remains to be discovered regarding the roles of protein phosphatases in mitogenic signaling pathways. The ability of okadaic acid to activate MAPK/ERKs demonstrates that alteration in serine/threonine dephosphorylation can have significant effects on common steps in growth stimulation induced by different types of mitogens. As in the case of cell cycle control, protein serine/threonine phosphatase plays a central role in the reentry of quiescent cells into the cycle. Because the only known targets of okadaic acid are the catalytic subunits PP1 and PP2A, these enzymes are crucial components of two basic functions carried out by cells: growth and division. Important and obligatory roles for PP2B, PP2C, and newly discovered serine/threonine phosphatases are also likely. However, the limited tissue distribution, unique regulatory properties, and limited substrate specificities of these forms suggest more specialized functions in restricted cell types. The available information on the specific functions of different forms of protein serine/threonine phosphatases, let alone their individual isoforms and different multimeric holoenzymes, is still severely limited. Years of biochemical characterization and cDNA cloning have left us with far more forms than functions. This has led to the gratifying situation, at least for the biochemists, in which genetics and cell biology identify protein phosphatases for which a wealth of biochemical information is already available. The appreciation of the importance of these enzymes in the coming years can only increase as the functions for individual forms are discovered.
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
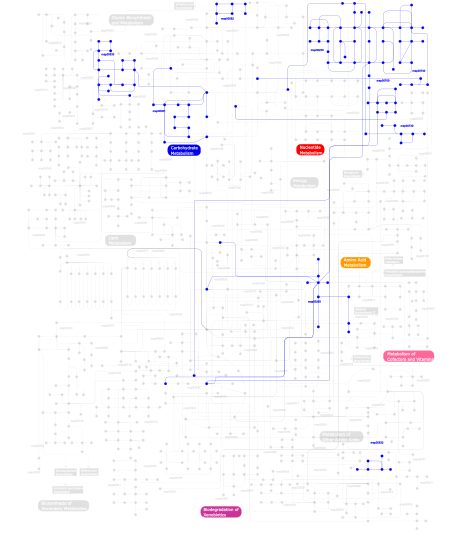
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 24.68 map04010 MAPK signaling pathway 10.39 map04115 p53 signaling pathway 10.39  map00230
map00230Purine metabolism 9.09 map04620 Toll-like receptor signaling pathway 6.49  map00760
map00760Nicotinate and nicotinamide metabolism 6.49  map00740
map00740Riboflavin metabolism 6.49  map00530
map00530Aminosugars metabolism 6.49  map00730
map00730Thiamine metabolism 6.49  map00051
map00051Fructose and mannose metabolism 5.19  map00562
map00562Inositol phosphate metabolism 5.19  map00632
map00632Benzoate degradation via CoA ligation 2.60  map00260
map00260Glycine, serine and threonine metabolism This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with PP2Cc domain which could be assigned to a KEGG orthologous group, and not all proteins containing PP2Cc domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of PP2Cc domains in PDB
PDB code Main view Title 1a6q 
CRYSTAL STRUCTURE OF THE PROTEIN SERINE/THREONINE PHOSPHATASE 2C AT 2 A RESOLUTION 1txo 
Crystal structure of the Mycobacterium tuberculosis serine/threonine phosphatase PstP/Ppp at 1.95 A. 2cm1 
Crystal structure of the catalytic domain of serine threonine protein phosphatase PstP in complex with 2 Manganese ions. 2i0o 
Crystal structure of Anopheles gambiae Ser/Thr phosphatase complexed with Zn2+ 2i44 
Crystal structure of serine-threonine phosphatase 2C from Toxoplasma gondii 2iq1 
Crystal structure of human PPM1K 2irm 
Crystal structure of mitogen-activated protein kinase kinase kinase 7 interacting protein 1 from Anopheles gambiae 2isn 
Crystal structure of a phosphatase from a pathogenic strain Toxoplasma gondii 2j4o 
Structure of TAB1 2j82 
Structural analysis of the PP2C Family Phosphatase tPphA from Thermosynechococcus elongatus 2j86 
Structural analysis of the PP2C Family Phosphatase tPphA of Thermosynechococcus elongatus 2jfr 
Crystal structure of the PP2C Ser-Thr phosphatase MsPP from Mycobacterium smegmatis in complex with phosphate at 0.83 A resolution 2jfs 
Crystal structure of the PPM Ser-Thr phosphatase MsPP from Mycobacterium smegmatis in complex with cacodylate 2jft 
Crystal structure of the PPM Ser-Thr phosphatase MsPP from Mycobacterium smegmatis in complex with sulfate 2p8e 
Crystal structure of the serine/threonine phosphatase domain of human PPM1B 2pk0 
Structure of the S. agalactiae serine/threonine phosphatase at 2.65 resolution 2pnq 
Crystal structure of pyruvate dehydrogenase phosphatase 1 (PDP1) 2pom 
TAB1 with manganese ion 2pop 
The Crystal Structure of TAB1 and BIR1 complex 2v06 
Crystal structure of the PPM Ser-Thr phosphatase MsPP from Mycobacterium smegmatis at pH 5.5 2xzv 
The cyanobacterial PP2C-like phosphatase tPphA requires three metals in the catalytic center for efficient catalysis 2y09 
The cyanobacterial PP2C-like phosphatase tPphA requires three metals in the catalytic center for efficient catalysis 3d8k 
Crsytal structure of a phosphatase from a toxoplasma gondii 3fxj 
Crystal Structure of Human Protein phosphatase 1A (PPM1A) Bound with Phosphate at 3 mM of Mn2+ 3fxk 
Crystal Structure of Human Protein phosphatase 1A (PPM1A) Bound with Phosphate at 10 mM of Mn2+ 3fxl 
Crystal Structure of Human Protein phosphatase 1A (PPM1A) Bound with Citrate at 1 mM of Mn2+ 3fxm 
Crystal Structure of Human Protein phosphatase 1A (PPM1A) Bound with Citrate at 10 mM of Mn2+ 3fxo 
Crystal Structure of Human Protein phosphatase 1A (PPM1A) Bound with Phosphate at 1 mM of Mn2+ 3jrq 
Crystal structure of (+)-ABA-bound PYL1 in complex with ABI1 3kb3 
Crystal structure of abscisic acid-bound PYL2 in complex with HAB1 3kdj 
Complex structure of (+)-ABA-bound PYL1 and ABI1 3mq3 
Crystal structure of native bovine PDP1c 3n3c 
Crystal structure of native bovine PDP1c 3nmn 
Crystal structure of pyrabactin-bound abscisic acid receptor PYL1 in complex with type 2C protein phosphatase ABI1 3nmt 
Crystal structure of pyrabactin bound abscisic acid receptor PYL2 mutant A93F in complex with type 2C protein phosphatase HAB1 3nmv 
Crystal structure of pyrabactin-bound abscisic acid receptor PYL2 mutant A93F in complex with type 2C protein phosphatase ABI2 3qn1 
Crystal structure of the PYR1 Abscisic Acid receptor in complex with the HAB1 type 2C phosphatase catalytic domain 3rnr 
Crystal Structure of Stage II Sporulation E Family Protein from Thermanaerovibrio acidaminovorans 3rt0 
Crystal structure of PYL10-HAB1 complex in the absence of abscisic acid (ABA) 3t91 
Structure of the Phosphatase Domain of the Cell Fate Determinant SpoIIE from Bacillus subtilis 3t9q 
Structure of the Phosphatase Domain of the Cell Fate Determinant SpoIIE from Bacillus subtilis (Mn presoaked) 3ujg 
Crystal structure of SnRK2.6 in complex with HAB1 3ujk 
Crystal structure of protein phosphatase ABI2 3ujl 
Crystal structure of abscisic acid bound PYL2 in complex with type 2C protein phosphatase ABI2 3w40 
Crystal structure of RsbX in complex with magnesium in space group P1 3w41 
Crystal structure of RsbX in complex with magnesium in space group P21 3w42 
Crystal structure of RsbX in complex with manganese in space group P1 3w43 
Crystal structure of RsbX in complex with manganese in space group P21 3w44 
Crystal structure of RsbX, selenomethionine derivative 3w45 
Crystal structure of RsbX in complex with cobalt in space group P1 3zt9 
The bacterial stressosome: a modular system that has been adapted to control secondary messenger signaling 3zvu 
Structure of the PYR1 His60Pro mutant in complex with the HAB1 phosphatase and Abscisic acid 4da1 
Crystal structure of branched-chain alpha-ketoacid dehydrogenase phosphatase with Mg (II) ions at the active site 4ds8 
Complex structure of abscisic acid receptor PYL3-(+)-ABA-HAB1 in the presence of Mn2+ 4jnd 
Structure of a C.elegans sex determining protein 4la7 
X-ray crystal structure of the PYL2-quinabactin-Hab1 ternary complex 4lg5 
ABA-mimicking ligand QUINABACTIN in complex with ABA receptor PYL2 and PP2C HAB1 4lga 
ABA-mimicking ligand N-(2-OXO-1-PROPYL-1,2,3,4-TETRAHYDROQUINOLIN-6-YL)-1-PHENYLMETHANESULFONAMIDE in complex with ABA receptor PYL2 and PP2C HAB1 4lgb 
ABA-mimicking ligand N-(1-METHYL-2-OXO-1,2,3,4-TETRAHYDROQUINOLIN-6-YL)-1-(4-METHYLPHENYL)METHANESULFONAMIDE in complex with ABA receptor PYL2 and PP2C HAB1 4n0g 
Crystal Structure of PYL13-PP2CA complex 4oic 
4OIC 4ra2 
4RA2 4raf 
4RAF 4rag 
4RAG 4wvo 
4WVO 4yzg 
4YZG 4yzh 
4YZH 5d2u 
5D2U 5f1m 
5F1M 5iti 
5ITI 5jo1 
5JO1 5jo2 
5JO2 - Links (links to other resources describing this domain)
-
PROSITE PP2C PFAM PP2C

