PQQbeta-propeller repeat |
|---|
| SMART accession number: | SM00564 |
|---|---|
| Description: | Beta-propeller repeat occurring in enzymes with pyrrolo-quinoline quinone (PQQ) as cofactor, in Ire1p-like Ser/Thr kinases, and in prokaryotic dehydrogenases. |
| Interpro abstract (IPR018391): | Pyrrolo-quinoline quinone (PQQ) is a redox coenzyme, which serves as a cofactor for a number of enzymes (quinoproteins) and particularly for some bacterial dehydrogenases [ (PUBMED:2549854) (PUBMED:2572081) (PUBMED:18838797) ]. A number of bacterial quinoproteins belong to this family. Enzymes in this group have repeats of a beta propeller. |
| Family alignment: |
There are 237300 PQQ domains in 48510 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing PQQ domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with PQQ domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing PQQ domain in the selected taxonomic class.
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Xia Z, Dai W, Zhang Y, White SA, Boyd GD, Mathews FS
- Determination of the gene sequence and the three-dimensional structure at 2.4 angstroms resolution of methanol dehydrogenase from Methylophilus W3A1.
- J Mol Biol. 1996; 259: 480-501
- Display abstract
The DNA sequences for the genes encoding the heavy and light subunits of methanol dehydrogenase from Methylophilus methylotrophus W3A1 have been determined. The deduced amino acid sequence has enabled the structure of the enzyme to be refined at 2.4 angstrom resolution against X-ray data collected on a Hamlin area detector. The structure was refined using the programs PROFFT and X-PLOR with several model building step interspersed. The final model contains two heavy chains (571 amino acids), two light chains (69 amino acids), two molecules of pyrroloquinoline quinone, two Ca2+ and 521 solvent molecules. Each half molecule contains four disulfide linkages and four cis peptides. One of the disulfides is formed from two adjacent cysteine residues linked by a trans peptide which creates a novel eight-membered ring. The heavy subunit is an 8-fold beta-propeller, each "blade" of which is a four-stranded antiparallel twisted beta-sheet. The light chain is an elongated subunit stretching across the surface of the heavy subunit, with residues 1 to 32 containing four beta-turns and residues 33 to 62 forming a helix; however, it neither interacts with the active site, nor the other HL dimer and its functional role is obscure. Around the 8-fold beta-propeller there is a repeating pattern of tryptophan residues located in the outer strand of seven of the eight beta-leaflets, each packed between adjacent leaflets. Each of these tryptophan residues is centered in the beta-strand and participates in the main chain hydrogen bonding of the sheet. Five of the seven tryptophan residues have closely similar interactions with the adjacent beta-leaflet including stacking of the tryptophan indole rings against a peptide plane and formation of a hydrogen bond from NE1 of the indole ring to a main-chain carbonyl. This repeating pattern is conserved over a number of MEDH sequences. The PQQ is located on the pseudo 8-fold rotation axis of the heavy subunit, in a funnel-shaped internal cavity, sandwiched between the indole ring of Trp237 and the two sulfur atoms of the Cys103-Cys104 vicinal disulfide. A hexacoordinate Ca2+ is bound in the active site by one nitrogen and five oxygen ligands, three from the PQQ and the others from two protein side-chains. In the active site an isolated solvent molecule is bound to the O5 of PQQ and to a nearby aspartate side-chain; its position may be the binding site for methanol. The aspartate might than serve as a general base for proton abstraction from the substrate hydroxyl. The C5 atom of PQQ could be activated by electrophilic catalysis by a nearby argenine side-chain or by the calcium ion bound to PQQ.
- Blake CC, Ghosh M, Harlos K, Avezoux A, Anthony C
- The active site of methanol dehydrogenase contains a disulphide bridge between adjacent cysteine residues.
- Nat Struct Biol. 1994; 1: 102-5
- Display abstract
Adjacent cysteine residues can only form disulphide bridges in a distorted structure containing a cis-peptide link. Such bridges are extremely uncommon, identified so far in the acetyl choline receptor alone where the structure of the bridge is undetermined. Here we present the first molecular description of a disulphide bridge of this type in the quinoprotein methanol dehydrogenase from Methylobacterium extorquens. We show that this structure occurs in close proximity to the pyrrolo-quinoline quinone prosthetic group and a calcium ion in the active site of the enzyme. This unusual disulphide bridge appears to play a role in the electron transfer reaction mediated by methanol dehydrogenase.
- Duine JA, Jongejan JA
- Quinoproteins, enzymes with pyrrolo-quinoline quinone as cofactor.
- Annu Rev Biochem. 1989; 58: 403-26
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
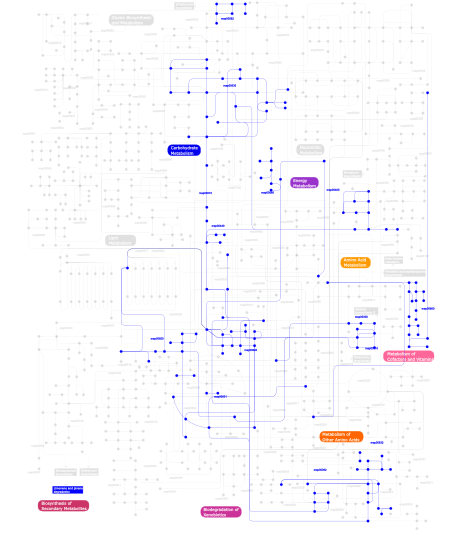
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 19.73  map00030
map00030Pentose phosphate pathway 13.87  map00680
map00680Methane metabolism 13.87  map00010
map00010Glycolysis / Gluconeogenesis 13.87  map00631
map006311,2-Dichloroethane degradation 13.87  map00650
map00650Butanoate metabolism 13.87  map00640
map00640Propanoate metabolism 2.13  map00440
map00440Aminophosphonate metabolism 2.13 map00903 Limonene and pinene degradation 2.13  map00362
map00362Benzoate degradation via hydroxylation 1.07  map00562
map00562Inositol phosphate metabolism 1.07  map00632
map00632Benzoate degradation via CoA ligation 1.07  map00400
map00400Phenylalanine, tyrosine and tryptophan biosynthesis 0.53 map03090 Type II secretion system 0.27  map00300
map00300Lysine biosynthesis 0.27  map00860
map00860Porphyrin and chlorophyll metabolism 0.27  map00310
map00310Lysine degradation This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with PQQ domain which could be assigned to a KEGG orthologous group, and not all proteins containing PQQ domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of PQQ domains in PDB
PDB code Main view Title 1flg 
CRYSTAL STRUCTURE OF THE QUINOPROTEIN ETHANOL DEHYDROGENASE FROM PSEUDOMONAS AERUGINOSA 1g72 
CATALYTIC MECHANISM OF QUINOPROTEIN METHANOL DEHYDROGENASE: A THEORETICAL AND X-RAY CRYSTALLOGRAPHIC INVESTIGATION 1h4i 
Methylobacterium extorquens methanol dehydrogenase 1h4j 
Methylobacterium extorquens methanol dehydrogenase D303E mutant 1kb0 
Crystal Structure of Quinohemoprotein Alcohol Dehydrogenase from Comamonas testosteroni 1kv9 
Structure at 1.9 A Resolution of a Quinohemoprotein Alcohol Dehydrogenase from Pseudomonas putida HK5 1lrw 
Crystal structure of methanol dehydrogenase from P. denitrificans 1w6s 
The high resolution structure of methanol dehydrogenase from methylobacterium extorquens 1yiq 
Molecular cloning and structural analysis of quinohemoprotein alcohol dehydrogenase ADHIIG from Pseudomonas putida HK5. Compariison to the other quinohemoprotein alcohol dehydrogenase ADHIIB found in the same microorganism. 2ad6 
crystal structure of methanol dehydrogenase from M. W3A1 (form C) 2ad7 
crystal structure of methanol dehydrogenase from M. W3A1 (form C) in the presence of methanol 2ad8 
crystal structure of methanol dehydrogenase from M. W3A1 (form C) in the presence of ethanol 2be1 
Structure of the compact lumenal domain of yeast Ire1 2d0v 
Crystal structure of methanol dehydrogenase from Hyphomicrobium denitrificans 2hz6 
The crystal structure of human IRE1-alpha luminal domain 2yh3 
The structure of BamB from E. coli 2yms 
Structure and assembly of a b-propeller with nine blades and a new conserved repetitive sequence motif 3hxj 
Crystal Structure of Pyrrolo-quinoline quinone (PQQ_DH) from Methanococcus maripaludis, Northeast Structural Genomics Consortium Target MrR86 3p1l 
Crystal structure of Escherichia coli BamB, a lipoprotein component of the beta-barrel assembly machinery complex, native crystals. 3prw 
Crystal structure of the lipoprotein BamB 3q54 
Crystal structure of Escherichia coli BamB 3q7m 
The crystal structure of BamB from the BAM complex in spacegroup I222 3q7n 
The crystal structure of BamB from the BAM complex in spacegroup P212121 3q7o 
The crystal structure of BamB from the BAM complex in spacegroup P213 4aah 
METHANOL DEHYDROGENASE FROM METHYLOPHILUS W3A1 4cvb 
4CVB 4cvc 
4CVC 4hdj 
Crystal Structure of BamB from Pseudomonas aeruginosa 4imm 
The crystal structure of BamB from Moraxella catarrhalis 4mae 
Methanol dehydrogenase from Methylacidiphilum fumariolicum SolV 4mh1 
4MH1 4pk1 
4PK1 4tqo 
4TQO 4xga 
4XGA 5ayw 
5AYW 5d0o 
5D0O 5ljo 
5LJO - Links (links to other resources describing this domain)
-
INTERPRO IPR018391

