The domain within your query sequence starts at position 1642 and ends at position 1703; the E-value for the SH3 domain shown below is 1.23e-7.
ELTVVLQDFSAGHSSELSIQVGQTVELLERPSERPGWCLVRTTERSPPQEGLVPSSALCI SH
SH3Src homology 3 domains |
|---|
| SMART accession number: | SM00326 |
|---|---|
| Description: | Src homology 3 (SH3) domains bind to target proteins through sequences containing proline and hydrophobic amino acids. Pro-containing polypeptides may bind to SH3 domains in 2 different binding orientations. |
| Interpro abstract (IPR001452): | SH3 (src Homology-3) domains are small protein modules containing approximately 50 amino acid residues [ (PUBMED:15335710) (PUBMED:11256992) ]. They are found in a great variety of intracellular or membrane-associated proteins [ (PUBMED:1639195) (PUBMED:14731533) (PUBMED:7531822) ] for example, in a variety of proteins with enzymatic activity, in adaptor proteins, such as fodrin and yeast actin binding protein ABP-1. The SH3 domain has a characteristic fold which consists of five or six beta-strands arranged as two tightly packed anti-parallel beta sheets. The linker regions may contain short helices. The surface of the SH3-domain bears a flat, hydrophobic ligand-binding pocket which consists of three shallow grooves defined by conservative aromatic residues in which the ligand adopts an extended left-handed helical arrangement. The ligand binds with low affinity but this may be enhanced by multiple interactions. The region bound by the SH3 domain is in all cases proline-rich and contains PXXP as a core-conserved binding motif. The function of the SH3 domain is not well understood but they may mediate many diverse processes such as increasing local concentration of proteins, altering their subcellular location and mediating the assembly of large multiprotein complexes [ (PUBMED:7953536) ]. The crystal structure of the SH3 domain of the cytoskeletal protein spectrin, and the solution structures of SH3 domains of phospholipase C (PLC-y) and phosphatidylinositol 3-kinase p85 alpha-subunit, have been determined [ (PUBMED:1279434) (PUBMED:7684655) (PUBMED:7681365) ]. In spite of relatively limited sequence similarity, their overall structures are similar. The domains belong to the alpha+beta structural class, with 5 to 8 beta-strands forming 2 tightly-packed, anti-parallel beta-sheets arranged in a barrel-like structure, and intervening loops sometimes forming helices. Conserved aliphatic and aromatic residues form a hydrophobic core (A11, L23, A29, V34, W42, L52 and V59 in PLC-y [ (PUBMED:7681365) ]) and a hydrophobic pocket on the molecular surface (L12, F13, W53 and P55 in PLC-y). The conserved core is believed to stabilise the fold, while the pocket is thought to serve as a binding site for target proteins. Conserved carboxylic amino acids located in the loops, on the periphery of the pocket (D14 and E22), may be involved in protein-protein interactions via proline-rich regions. The N- and C-terminal are packed in close proximity, indicating that they are independent structural modules. |
| GO function: | protein binding (GO:0005515) |
| Family alignment: |
There are 197921 SH3 domains in 149315 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing SH3 domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with SH3 domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing SH3 domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Cellular role: signalling
Binding / catalysis: protein-binding, proline-rich peptide-binding - Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Arold S et al.
- The crystal structure of HIV-1 Nef protein bound to the Fyn kinase SH3 domain suggests a role for this complex in altered T cell receptor signaling.
- Structure. 1997; 5: 1361-72
- Display abstract
BACKGROUND: Human immunodeficiency virus (HIV) Nef protein accelerates virulent progression of acquired immunodeficiency syndrome (AIDS) by its interaction with specific cellular proteins involved in signal transduction and host cell activation. Nef has been shown to bind specifically to a subset of the Src family of kinases. The structures of free Nef and Nef bound to Src homology region 3 (SH3) domain are important for the elucidation of how the affinity and specificity for the Src kinase family SH3 domains are achieved, and also for the development of potential drugs and vaccines against AIDS. RESULTS: We have determined the crystal structures of the conserved core of HIV-1 Nef protein alone and in complex with the wild-type SH3 domain of the p59fyn protein tyrosine kinase (Fyn), at 3.0 A resolution. Comparison of the bound and unbound Nef structures revealed that a proline-rich motif (Pro-x-x-Pro), which is implicated in SH3 binding, is partially disordered in the absence of the binding partner; this motif only fully adopts a left-handed polyproline type II helix conformation upon complex formation with the Fyn SH3 domain. In addition, the structures show how an arginine residue (Arg77) of Nef interacts with Asp 100 of the so-called RT loop within the Fyn SH3 domain, and triggers a hydrogen-bond rearrangement which allows the loop to adapt to complement the Nef surface. The Arg96 residue of the Fyn SH3 domain is specifically accommodated in the same hydrophobic pocket of Nef as the isoleucine residue of a previously described Fyn SH3 (Arg96-->lle) mutant that binds to Nef with higher affinity than the wild type. CONCLUSIONS: The three-dimensional structures support evidence that the Nef-Fyn complex forms in vivo and may have a crucial role in the T cell perturbating action of Nef by altering T cell receptor signaling. The structures of bound and unbound Nef reveal that the multivalency of SH3 binding may be achieved by a ligand induced flexibility in the RT loop. The structures suggest possible targets for the design of inhibitors which specifically block Nef-SH3 interactions.
- Barrett T et al.
- The structure of the GTPase-activating domain from p50rhoGAP.
- Nature. 1997; 385: 458-61
- Display abstract
Members of the Rho family of small G proteins transduce signals from plasma-membrane receptors and control cell adhesion, motility and shape by actin cytoskeleton formation. They also activate other kinase cascades. Like all other GTPases, Rho proteins act as molecular switches, with an active GTP-bound form and an inactive GDP-bound form. The active conformation is promoted by guanine-nucleotide exchange factors, and the inactive state by GTPase-activating proteins (GAPs) which stimulate the intrinsic GTPase activity of small G proteins. Rho-specific GAP domains are found in a wide variety of large, multi-functional proteins. Here we report the crystal structure of an active 242-residue C-terminal fragment of human p50rhoGAP. The structure is an unusual arrangement of nine alpha-helices, the core of which includes a four-helix bundle. Residues conserved across the rhoGAP family are largely confined to one face of this bundle, which may be an interaction site for target G proteins. In particular, we propose that Arg 85 and Asn 194 are involved in binding G proteins and enhancing GTPase activity.
- Cowburn D
- Peptide recognition by PTB and PDZ domains.
- Curr Opin Struct Biol. 1997; 7: 835-8
- Display abstract
Protein tyrosine binding (PTB) and 'post synaptic density disc-large zo-1' (PDZ) domains bind to short peptidic ligands by augmentation of one of the domain's beta sheets and other recognition mechanisms. The two domain classes have a superficial resemblance to each other, even though no sequential homology exists. The structural bases of the interactions are well understood for the few domains now experimentally determined, and ligand-target pairs can probably be identified in favorable cases by analogy with the known domains. For both PTB and PDZ classes, functional activities are still not fully defined: it is possible that these domain classes, along with pleckstrin homology domains, have multiple roles.
- Han L et al.
- Protein binding and signaling properties of RIN1 suggest a unique effector function.
- Proc Natl Acad Sci U S A. 1997; 94: 4954-9
- Display abstract
Human RIN1 was first characterized as a RAS binding protein based on the properties of its carboxyl-terminal domain. We now show that full-length RIN1 interacts with activated RAS in mammalian cells and defines a minimum region of 434 aa required for efficient RAS binding. RIN1 interacts with the "effector domain" of RAS and employs some RAS determinants that are common to, and others that are distinct from, those required for the binding of RAF1, a known RAS effector. The same domain of RIN1 that binds RAS also interacts with 14-3-3 proteins, extending the similarity between RIN1 and other RAS effectors. When expressed in mammalian cells, the RAS binding domain of RIN1 can act as a dominant negative signal transduction blocker. The amino-terminal domain of RIN1 contains a proline-rich sequence similar to consensus Src homology 3 (SH3) binding regions. This RIN1 sequence shows preferential binding to the ABL-SH3 domain in vitro. Moreover, the amino-terminal domain of RIN1 directly associates with, and is tyrosine phosphorylated by, c-ABL. In addition, RIN1 encodes a functional SH2 domain that has the potential to activate downstream signals. These data suggest that RIN1 is able to mediate multiple signals. A differential pattern of expression and alternate splicing indicate several levels of RIN1 regulation.
- Hildebrandt F et al.
- A novel gene encoding an SH3 domain protein is mutated in nephronophthisis type 1.
- Nat Genet. 1997; 17: 149-53
- Display abstract
Juvenile nephronophthisis (NPH), an autosomal recessive cystic kidney disease, is the primary genetic cause of chronic renal failure in children. About two thirds of patients with NPH carry a large homozygous deletion at the gene locus NPH1 on 2q13. We here identify a novel gene. NPHP1, which extends over most of this common deletion. The 4.5-kb transcript encodes a protein with an SH3 domain, which is highly conserved throughout evolution. The 11-kb interval between the 3' end of NPHP1 and an inverted repeat containing the distal deletion breakpoint was found to contain the first exon of a second gene, MALL. In patients with a hemizygous deletion of the NPH1 region, additional point mutations were found in NPHP1 but not in MALL.
- Kuriyan J, Cowburn D
- Modular peptide recognition domains in eukaryotic signaling.
- Annu Rev Biophys Biomol Struct. 1997; 26: 259-88
- Display abstract
A characteristic feature of cellular signal transduction pathways in eukaryotes is the separation of catalysis from target recognition. Several modular domains that recognize short peptide sequences and target signaling proteins to these sequences have been identified. The structural bases of the specificities of recognition by SH2, SH3, and PTB domains have been elucidated by X-ray crystallography and NMR, and these results are reviewed here. In addition, the mechanism of cooperative interactions between these domains is discussed.
- Sicheri F, Moarefi I, Kuriyan J
- Crystal structure of the Src family tyrosine kinase Hck.
- Nature. 1997; 385: 602-9
- Display abstract
The crystal structure of the haematopoietic cell kinase Hck has been determined at 2.6/2.9 A resolution. Inhibition of enzymatic activity is a consequence of intramolecular interactions of the enzyme's Src-homology domains SH2 and SH3, with concomitant displacement of elements of the catalytic domain. The conformation of the active site has similarities with that of inactive cyclin-dependent protein kinases.
- Xu W, Harrison SC, Eck MJ
- Three-dimensional structure of the tyrosine kinase c-Src.
- Nature. 1997; 385: 595-602
- Display abstract
The structure of a large fragment of the c-Src tyrosine kinase, comprising the regulatory and kinase domains and the carboxy-terminal tall, has been determined at 1.7 A resolution in a closed, inactive state. Interactions among domains, stabilized by binding of the phosphorylated tail to the SH2 domain, lock the molecule in a conformation that simultaneously disrupts the kinase active site and sequesters the binding surfaces of the SH2 and SH3 domains. The structure shows how appropriate cellular signals, or transforming mutations in v-Src, could break these interactions to produce an open, active kinase.
- Lee CH, Saksela K, Mirza UA, Chait BT, Kuriyan J
- Crystal structure of the conserved core of HIV-1 Nef complexed with a Src family SH3 domain.
- Cell. 1996; 85: 931-42
- Display abstract
The crystal structure of the conserved core of HIV-1 Nef has been determined in complex with the SH3 domain of a mutant Fyn tyrosine kinase (a single amino acid substitution, Arg-96 to isoleucine), to which Nef binds tightly. The conserved PxxP sequence motif of Nef, known to be important for optimal viral replication, is part of a polyproline type II helix that engages the SH3 domain in a manner resembling closely the interaction of isolated peptides with SH3 domains. The Nef-SH3 structure also reveals how high affinity and specificity in the SH3 interaction is achieved by the presentation of the PxxP motif within the context of the folded structure of Nef.
- Metzler WJ, Leiting B, Pryor K, Mueller L, FarmerBT2nd
- The three-dimensional solution structure of the SH2 domain from p55blk kinase.
- Biochemistry. 1996; 35: 6201-11
- Display abstract
Signal transduction in B cells is mediated, in part, by the interaction of the cytoplasmic components of the antigen receptor complex and various members of the src family tyrosine kinases. Key to this process appears to be the interaction of the tyrosine kinase SH2 domains with the tyrosine-phosphorylated cytoplasmic domain of Ig-alpha, a disulfide-bonded heterodimeric (with Ig-beta or Ig-gamma) transmembrane protein that noncovalently associates with the antigen receptor immunoglobin chains. In addition to binding to the phosphorylated cytoplasmic domains of Ig-alpha and Ig-beta, blk and fyn(T), two members of the src family kinases, have been shown to bind overlapping but distinct sets of phosphoproteins [Malek & Desiderio (1993) J. Biol. Chem. 268. 22557-22565]. A comparison of their three-dimensional structures may elucidate the apparently subtle differences required for phosphoprotein discrimination. To begin characterizing the blk/fyn/phosphosphoprotein interactions, we have determined the three-dimensional solution structure of the SH2 domain of blk kinase by nuclear magnetic resonance (NMR) spectroscopy. 1H, 13C, and 15N resonances of the SH2 domain of blk kinase were assigned by analysis of multidimensional, double- and triple-resonance NMR experiments. Twenty structures of the blk SH2 domain were refined with the program X-PLOR using a total of 2080 experimentally derived conformational restraints. The structures converged to a root-mean-squared (rms) distance deviation of 0.51 and 0.95 A for the backbone atoms and for the non-hydrogen atoms, respectively. The blk SH2 domain adopts the prototypical SH2 fold. Structurally, blk SH2 is most similar to the crystal structure of the v-src SH2 domain [Waksman et al. (1993) Nature 358.646-653] and superimposes on the crystal structure with an rmsd of 1.52 A for the backbone atoms. The largest deviations occur in the four loops interconnecting beta-strands A-E, which are the least well-defined regions in the NMR structure. Exclusion of these loops lowers this rmsd to 0.82 A. The conformation of the BC loop in the blk SH2 domain is similar to the open conformation in the apo lck SH2 domain, suggesting that, like the lck SH2 domain, the blk SH2 domain may have a gated phosphopeptide binding site. Finally, it is proposed that the amino acid substitution of Lys 88 (blk) for Glu [fyn(T)] is important for the observed differences in specificity between blk and fyn(T) SH2 domains.
- Mott HR, Carpenter JW, Zhong S, Ghosh S, Bell RM, Campbell SL
- The solution structure of the Raf-1 cysteine-rich domain: a novel ras and phospholipid binding site.
- Proc Natl Acad Sci U S A. 1996; 93: 8312-7
- Display abstract
The Raf-1 protein kinase is the best-characterized downstream effector of activated Ras. Interaction with Ras leads to Raf-1 activation and results in transduction of cell growth and differentiation signals. The details of Raf-1 activation are unclear, but our characterization of a second Ras-binding site in the cysteine-rich domain (CRD) and the involvement of both Ras-binding sites in effective Raf-1-mediated transformation provides insight into the molecular aspects and consequences of Ras-Raf interactions. The Raf-1 CRD is a member of an emerging family of domains, many of which are found within signal transducing proteins. Several contain binding sites for diacylglycerol (or phorbol esters) and phosphatidylserine and are believed to play a role in membrane translocation and enzyme activation. The CRD from Raf-1 does not bind diacylglycerol but interacts with Ras and phosphatidylserine. To investigate the ligand-binding specificities associated with CRDs, we have determined the solution structure of the Raf-1 CRD using heteronuclear multidimensional NMR. We show that there are differences between this structure and the structures of two related domains from protein kinase C (PKC). The differences are confined to regions of the CRDs involved in binding phorbol ester in the PKC domains. Since phosphatidylserine is a common ligand, we expect its binding site to be located in regions where the structures of the Raf-1 and PKC domains are similar. The structure of the Raf-1 CRD represents an example of this family of domains that does not bind diacylglycerol and provides a framework for investigating its interactions with other molecules.
- Muchmore SW et al.
- X-ray and NMR structure of human Bcl-xL, an inhibitor of programmed cell death.
- Nature. 1996; 381: 335-41
- Display abstract
THE Bcl-2 family of proteins regulate programmed cell death by an unknown mechanism. Here we describe the crystal and solution structures of a Bcl-2 family member, Bcl-xL (ref. 2). The structures consist of two central, primarily hydrophobic alpha-helices, which are surrounded by amphipathic helices. A 60-residue loop connecting helices alpha1 and alpha2 was found to be flexible and non-essential for anti-apoptotic activity. The three functionally important Bcl-2 homology regions (BH1, BH2 and BH3) are in close spatial proximity and form an elongated hydrophobic cleft that may represent the binding site for other Bcl-2 family members. The arrangement of the alpha-helices in Bcl-xL is reminiscent of the membrane translocation domain of bacterial toxins, in particular diphtheria toxin and the colicins. The structural similarity may provide a clue to the mechanism of action of the Bcl-2 family of proteins.
- Musacchio A, Cantley LC, Harrison SC
- Crystal structure of the breakpoint cluster region-homology domain from phosphoinositide 3-kinase p85 alpha subunit.
- Proc Natl Acad Sci U S A. 1996; 93: 14373-8
- Display abstract
Proteins such as the product of the break-point cluster region, chimaerin, and the Src homology 3-binding protein 3BP1, are GTPase activating proteins (GAPs) for members of the Rho subfamily of small GTP-binding proteins (G proteins or GTPases). A 200-residue region, named the breakpoint cluster region-homology (BH) domain, is responsible for the GAP activity. We describe here the crystal structure of the BH domain from the p85 subunit of phosphatidylinositol 3-kinase at 2.0 A resolution. The domain is composed of seven helices, having a previously unobserved arrangement. A core of four helices contains most residues that are conserved in the BH family. Their packing suggests the location of a G-protein binding site. This structure of a GAP-like domain for small GTP-binding proteins provides a framework for analyzing the function of this class of molecules.
- Sparks AB, Hoffman NG, McConnell SJ, Fowlkes DM, Kay BK
- Cloning of ligand targets: systematic isolation of SH3 domain-containing proteins.
- Nat Biotechnol. 1996; 14: 741-4
- Display abstract
Based on the prevalence of modular protein domains, such as Src homology domain 3 and 2 (SH3 and SH2), among important signaling molecules, we have sought to identify new SH3 domain-containing proteins. However, modest sequence similarity among these domains restricts the use of DNA-based methods for this purpose. To circumvent this limitation, we have developed a functional screen that permits the rapid cloning of modular domains based on their ligand-binding activity. Using operationally defined SH3 ligands from combinatorial peptide libraries, we screened a series of mouse and human cDNA expression libraries. We found that 69 of the 74 clones isolated encode at least one SH3 domain. These clones encode 18 different SH3-containing proteins, 10 of which have not been described previously. The isolation of entire repertoires of modular domain-containing proteins will prove invaluable in genome analysis and in bringing new targets into drug discovery programs.
- Thornton KH, Mueller WT, McConnell P, Zhu G, Saltiel AR, Thanabal V
- Nuclear magnetic resonance solution structure of the growth factor receptor-bound protein 2 Src homology 2 domain.
- Biochemistry. 1996; 35: 11852-64
- Display abstract
A family of NMR solution structures of the growth factor receptor-bound protein 2 (Grb2) SH2 domain has been determined by heteronuclear multidimensional NMR. Proton, nitrogen, and carbon chemical shift assignments have been made for the SH2 domain of Grb2. Assignments were made from a combination of homonuclear two-dimensional and 15N- and 13C-edited three-dimensional spectra at pH 6.2 and 298 K. Structure-induced proton and carbon secondary shifts were calculated and used to facilitate the spectral assignment process. NOE, scalar coupling, secondary chemical shift, and amide proton exchange data were used to characterize the secondary structural elements and hydrogen-bonding network in the Grb2 SH2 domain. The three-dimensional structure of the Grb2 SH2 domain was calculated using 1112 restraints obtained from NOE, coupling constant, and amide proton exchange data. The rmsd for the 24 calculated structures to the mean structure of the Grb2 SH2 domain was 0.75 A for backbone and 1.28 A for all heavy atoms. The three-dimensional fold of the Grb2 SH2 domain is similar to that observed for other SH2 domains and consists of two alpha-helical segments and eight beta-strands, six strands that make up two contiguous antiparallel beta-sheets, and two strands that form two short parallel beta-sheets. The structure of the phosphotyrosine binding pocket of Grb2 is similar to that observed for other SH2 domains. The hydrophobic binding pocket of Grb2 is similar to that observed for Src with the exception that tryptophan 121 of Grb2 occupies part of the pY+3 binding pocket. Structural implications for the Grb2 SH2 domain selectivity at the pY+2 and pY+3 sites are discussed.
- Feng S, Kasahara C, Rickles RJ, Schreiber SL
- Specific interactions outside the proline-rich core of two classes of Src homology 3 ligands.
- Proc Natl Acad Sci U S A. 1995; 92: 12408-15
- Display abstract
Two dodecapeptides belonging to distinct classes of Src homology 3 (SH3) ligands and selected from biased phage display libraries were used to investigate interactions between a specificity pocket in the Src SH3 domain and ligant residues flanking the proline-rich core. The solution structures of c-Src SH3 complexed with these peptides were solved by NMR. In addition to proline-rich, polyproline type II helix-forming core, the class I and II ligands each possesses a flanking sequence that occupies a large pocket between the RT and n-Src loops of the SH3 domain. Structural and mutational analyses illustrate how the two classes of SH3 ligands exploit a specificity pocket on the receptor differently to increase binding affinity and specificity.
- Maignan S, Guilloteau JP, Fromage N, Arnoux B, Becquart J, Ducruix A
- Crystal structure of the mammalian Grb2 adaptor.
- Science. 1995; 268: 291-3
- Display abstract
The mammalian growth factor receptor-binding protein Grb2 is an adaptor that mediates activation of guanine nucleotide exchange on Ras. Grb2 binds to the receptor through its SH2 domain and to the carboxyl-terminal domain of Son of sevenless through its two SH3 domains. It is thus a key element in the signal transduction pathway. The crystal structure of Grb2 was determined to 3.1 angstrom resolution. The asymmetric unit is composed of an embedded dimer. The interlaced junctions between the SH2 and SH3 domains bring the two adjacent faces of the SH3 domains in van der Waals contact but leave room for the binding of proline-rich peptides.
- Mikol V, Baumann G, Zurini MG, Hommel U
- Crystal structure of the SH2 domain from the adaptor protein SHC: a model for peptide binding based on X-ray and NMR data.
- J Mol Biol. 1995; 254: 86-95
- Display abstract
Src homology 2 domains (SH2) are protein molecules found within a wide variety of cytoplasmic signalling molecules that bind with high affinity to phosphotyrosyl (pY)-containing protein sequences. We report here for crystal structure of the SH2 domain from the adaptor protein SHC (Shc), which has been refined by restrained least-squares methods to an R-factor of 17.3% to 2.7 A. The overall Shc architecture is essentially similar to that determined in other SH2 domains but it shows significant differences in a number of loops, thus providing a molecular surface with no obvious secondary pocket. Based on the knowledge of the crystal structure of the protein a model for a low affinity Shc-bound peptide has been generated from nuclear magnetic resonance data in solution using transferred nuclear Overhauser enhancements as intramolecular distance restraints. The model shows that the tyrosine moiety binds Shc in a rather similar way to that observed for other SH2-peptide complexes, but that the residue in position +3 does not seem to make specific contact with the protein. An intermolecular crystallographic interaction occurs between the pY-binding site and the C-terminal residues of a symmetry-related molecule. This crystal packing interaction suggests how inhibitory regulation could play a role in SHC activity.
- Narula SS et al.
- Solution structure of the C-terminal SH2 domain of the human tyrosine kinase Syk complexed with a phosphotyrosine pentapeptide.
- Structure. 1995; 3: 1061-73
- Display abstract
BACKGROUND: Recruitment of the intracellular tyrosine kinase Syk to activated immune-response receptors is a critical early step in intracellular signaling. In mast cells, Syk specifically associates with doubly phosphorylated immunoreceptor tyrosine-based activation motifs (ITAMs) that are found within the IgE receptor. The mechanism by which Syk recognizes these motifs is not fully understood. Both Syk SH2 (Src homology 2) domains are required for high-affinity binding to these motifs, but the C-terminal SH2 domain (Syk-C) can function independently and can bind, in isolation, to the tyrosine-phosphorylated IgE receptor in vitro. In order to improve understanding of the cellular function of Syk, we have determined the solution structure of Syk-C complexed with a phosphotyrosine peptide derived from the gamma subunit of the IgE receptor. RESULTS: The Syk-C:peptide structure is compared with liganded structures of both the SH2 domain of Src and the C-terminal SH2 domain of ZAP-70 (the 70 kDa zeta-associated protein). The topologies of these domains are similar, although significant differences occur in the loop regions. In the Syk-C structure, the phosphotyrosine and leucine residues of the peptide ligand interact with pockets on the protein, and the intervening residues are extended. CONCLUSIONS: Syk-C resembles other SH2 domains in its peptide-binding interactions and overall topology, a result that is consistent with its ability to function as an independent SH2 domain in vitro. This result suggests that Syk-C plays a unique role in the intact Syk protein. The determinants of the binding affinity and selectivity of Syk-C may reside in the least-conserved structural elements that comprise the phosphotyrosine- and leucine-binding sites. These structural features can be exploited for the design of Syk-selective SH2 antagonists for the treatment of allergic disorders and asthma.
- Trub T et al.
- Specificity of the PTB domain of Shc for beta turn-forming pentapeptide motifs amino-terminal to phosphotyrosine.
- J Biol Chem. 1995; 270: 18205-8
- Display abstract
Shc phosphorylation in cells following growth factor, insulin, cytokine, and lymphocyte receptor activation leads to its association with Grb2 and activation of Ras. In addition to being a cytoplasmic substrate of tyrosine kinases, Shc contains an SH2 domain and a non-SH2 phosphotyrosine binding (PTB) domain. Here we show that the Shc PTB domain, but not the SH2 domain, binds with high affinity (ID50 approximately equal to 1 microM) to phosphopeptides corresponding to the sequence surrounding Tyr250 of the polyoma virus middle T (mT) antigen (LLSNPTpYSVMRSK). Truncation studies show that five residues amino-terminal to tyrosine are required for high affinity binding, whereas all residues carboxyl-terminal to tyrosine can be deleted without loss of affinity. Substitution studies show that tyrosine phosphorylation is required and residues at -5, -3, -2, and -1 positions relative to pTyr are important for this interaction. 1H NMR studies demonstrate that the phosphorylated mT antigen-derived sequence forms a stable beta turn in solution, and correlations between structure and function indicate that the beta turn is important for PTB domain recognition. These results show that PTB domains are functionally distinct from SH2 domains. Whereas SH2 domain binding specificity derives from peptide sequences carboxyl-terminal to phosphotyrosine, the Shc PTB domain gains specificity by interacting with beta turn-forming sequences amino-terminal to phosphotyrosine.
- Xu RX, Word JM, Davis DG, Rink MJ, WillardDHJ r, GampeRTJ r
- Solution structure of the human pp60c-src SH2 domain complexed with a phosphorylated tyrosine pentapeptide.
- Biochemistry. 1995; 34: 2107-21
- Display abstract
Human pp60c-src is a cellular nonreceptor tyrosine kinase that participates in cytosolic signal transduction and has been implicated in the development of malignant tumors in the human breast and colon. Signal transduction is mediated by highly specific interactions between the SH2 domain and receptor phosphorylated tyrosine binding motifs. To elucidate the molecular conformation and interactions in solution, a family of highly resolved nuclear magnetic resonance (NMR) structures was determined for the src SH2 domain complexed with a high-affinity phosphorylated pentapeptide, acetyl-p YEEIE-OH. The 23 structures, generated with a distance geometry (DG) and a dynamical simulated annealing (SA) procedure, satisfied 2072 experimental restraints derived from a variety of multifrequency/multidimensional and isotope-filtered NMR data. Superimposition of residues 143-245 upon the mean coordinate set yielded an atomic rmsd of 0.58 +/- 0.09 A for the N, C alpha, C' atoms and 1.04 +/- 0.08 for all the non-hydrogen atoms. Residues in the ordered secondary structure regions superimpose to 0.29 +/- 0.04 A for the N, C alpha, C' and 0.73 +/- 0.08 A for all the non-hydrogen atoms. The angular order parameter calculated for the phi, psi angles was > 0.9 for 81 of the 106 protein residues. The main protein conformational features are three antiparallel beta-strands that traverse a compact core with an alpha-helix on each side of the core near the N- and C-termini. The observed intermolecular nuclear Overhauser effects (NOE) from the pY, +1E, and +3I residues positioned the ligand in an extended conformation across the SH2 domain surface with the pY and +3I side chains inserted into the protein binding pockets. In general, the protein conformation is consistent with previously reported structures of different SH2 domain complexes determined by X-ray crystallography. However, inter- or intramolecular interactions involving the guanidinium side chains of the solvated R alpha A2 or the buried R beta B5 were not observed at pH = 5.5 or 7.0. If such interactions exist in solution, the absence of any confirming data probably arises from rapid exchange with solvent and/or undetermined dynamic components. Thus, the unrestrained R alpha A2 side chain did not show an amino-aromatic interaction or a hydrogen bond to the -1 carbonyl oxygen as observed in the crystal structures. This result is consistent with the solution structure of a different SH2 domain complex. A more detailed comparison between the crystal structure and the NMR-derived solution structures of the same src SH2 domain complex is presented.(ABSTRACT TRUNCATED AT 400 WORDS)
- Borchert TV, Mathieu M, Zeelen JP, Courtneidge SA, Wierenga RK
- The crystal structure of human CskSH3: structural diversity near the RT-Src and n-Src loop.
- FEBS Lett. 1994; 341: 79-85
- Display abstract
SH3 domains are modules occurring in diverse proteins, ranging from cytoskeletal proteins to signaling proteins, such as tyrosine kinases. The crystal structure of the SH3 domain of Csk (c-Src specific tyrosine kinase) has been refined at a resolution of 2.5 A, with an R-factor of 22.4%. The structure is very similar to the FynSH3 crystal structure. When comparing CskSH3 and FynSH3 it is seen that the structural and charge differences of the RT-Src loop and the n-Src loop, near the conserved Trp47, correlate with different binding properties of these SH3 domains. The structure comparison suggests that those glycines and acid residues which are very well conserved in the SH3 sequences are important for the stability of the SH3 fold.
- Eck MJ, Atwell SK, Shoelson SE, Harrison SC
- Structure of the regulatory domains of the Src-family tyrosine kinase Lck.
- Nature. 1994; 368: 764-9
- Display abstract
The kinase p56lck (Lck) is a T-lymphocyte-specific member of the Src family of non-receptor tyrosine kinases. Members of the Src family each contain unique amino-terminal regions, followed by Src-homology domains SH3 and SH2, and a tyrosine kinase domain. SH3 and SH2 domains mediate critical protein interactions in many signal-transducing pathways. They are small, independently folded modules of about 60 and 100 residues, respectively, and they are often but not always found together in the same molecule. Like all nine Src-family kinases (reviewed in ref. 3), Lck is regulated by phosphorylation of a tyrosine in the short C-terminal tail of its catalytic domain. There is evidence that binding of the phosphorylated tail to the SH2 domain inhibits catalytic activity of the kinase domain and that the SH3 and SH2 domains may act together to effect this regulation. Here we report the crystal structures for a fragment of Lck bearing its SH3 and SH2 domains, alone and in complex with a phosphotyrosyl peptide containing the sequence of the Lck C-terminal regulatory tail. The latter complex represents the regulatory apparatus of Lck. The SH3-SH2 fragment forms similar dimers in both crystals, and the tail peptide binds at the intermolecular SH3/SH2 contact. The two structures show how an SH3 domain might recognize a specific target and suggest how dimerization could play a role in regulating Src-family kinases.
- Feng S, Chen JK, Yu H, Simon JA, Schreiber SL
- Two binding orientations for peptides to the Src SH3 domain: development of a general model for SH3-ligand interactions.
- Science. 1994; 266: 1241-7
- Display abstract
Solution structures of two Src homology 3 (SH3) domain-ligand complexes have been determined by nuclear magnetic resonance. Each complex consists of the SH3 domain and a nine-residue proline-rich peptide selected from a large library of ligands prepared by combinatorial synthesis. The bound ligands adopt a left-handed polyproline type II (PPII) helix, although the amino to carboxyl directionalities of their helices are opposite. The peptide orientation is determined by a salt bridge formed by the terminal arginine residues of the ligands and the conserved aspartate-99 of the SH3 domain. Residues at positions 3, 4, 6, and 7 of both peptides also intercalate into the ligand-binding site; however, the respective proline and nonproline residues show exchanged binding positions in the two complexes. These structural results led to a model for the interactions of SH3 domains with proline-rich peptides that can be used to predict critical residues in complexes of unknown structure. The model was used to identify correctly both the binding orientation and the contact and noncontact residues of a peptide derived from the nucleotide exchange factor Sos in association with the amino-terminal SH3 domain of the adaptor protein Grb2.
- Lim WA, Richards FM, Fox RO
- Structural determinants of peptide-binding orientation and of sequence specificity in SH3 domains.
- Nature. 1994; 372: 375-9
- Display abstract
The Src-homology-3 (SH3) domains of the Caenorhabditis elegans protein SEM-5 and its human and Drosophila homologues, Grb2 and Drk (refs 1-4), bind proline-rich sequences found in the nucleotide-exchange factor Sos as part of their proposed function linking receptor tyrosine kinase activation to Ras activation. Here we report the crystal structure at 2.0 A resolution of the carboxy-terminal SH3 domain from SEM-5 complexed to the mSos-derived amino-acid sequence PPPVPPRRR. The peptide is found to bind in an orientation ('minus') that is precisely opposite to that observed previously ('plus' orientation) in other SH3-peptide complexes. This novel ability of peptide-recognition proteins to recognize peptides in two distinct modes may play an important role in the signalling specificity of pathways involving SH3 domains. Comparison of this structure with other SH3 complexes reveals how a conserved binding face can be used to recognize peptides in different orientations, and why the Sos peptide binds in this particular orientation.
- Musacchio A, Saraste M, Wilmanns M
- High-resolution crystal structures of tyrosine kinase SH3 domains complexed with proline-rich peptides.
- Nat Struct Biol. 1994; 1: 546-51
- Display abstract
Src-homology 3 (SH3) domains bind to proline-rich motifs in target proteins. We have determined high-resolution crystal structures of the complexes between the SH3 domains of Abl and Fyn tyrosine kinases, and two ten-residue proline-rich peptides derived from the SH3-binding proteins 3BP-1 and 3BP-2. The X-ray data show that the basic mode of binding of both proline-rich peptides is the same. Peptides are bound over their entire length and interact with three major sites on the SH3 molecules by both hydrogen-bonding and van der Waals contacts. Residues 4-10 of the peptide adopt the conformation of a left-handed polyproline helix type II. Binding of the proline at position 2 requires a kink at the non-proline position 3.
- Perez-Alvarado GC et al.
- Structure of the carboxy-terminal LIM domain from the cysteine rich protein CRP.
- Nat Struct Biol. 1994; 1: 388-98
- Display abstract
The three dimensional solution structure of the carboxy terminal LIM domain of the avian Cysteine Rich Protein (CRP) has been determined by nuclear magnetic resonance spectroscopy. The domain contains two zinc atoms bound independently in CCHC (C = Cys, H = His) and CCCC modules. Both modules contain two orthogonally-arranged antiparallel beta-sheets, and the CCCC module contains an alpha-helix at its C terminus. The modules pack due to hydrophobic interactions forming a novel global fold. The structure of the C-terminal CCCC module is essentially identical to that observed for the DNA-interactive CCCC modules of the GATA-1 and steroid hormone receptor DNA binding domains, raising the possibility that the LIM motif may have a DNA binding function.
- Wittekind M et al.
- Orientation of peptide fragments from Sos proteins bound to the N-terminal SH3 domain of Grb2 determined by NMR spectroscopy.
- Biochemistry. 1994; 33: 13531-9
- Display abstract
NMR spectroscopy has been used to characterize the protein-protein interactions between the mouse Grb2 (mGrb2) N-terminal SH3 domain complexed with a 15-residue peptide (SPLLPKLPP-KTYKRE) corresponding to residues 1264-1278 of the mouse Sos-2 (mSos-2) protein. Intermolecular interactions between the peptide and 13C-15N-labeled SH3 domain were identified in half-reverse-filtered 2D and 3D NOESY experiments. Assignments for the protons involved in interactions between the peptide and the SH3 domain were confirmed in a series of NOESY experiments using a set of peptides in which different leucine positions were fully deuterated. The peptide ligand-binding site of the mGrb2 N-terminal SH3 domain is defined by the side chains of specific aromatic residues (Tyr7, Phe9, Trp36, Tyr52) that form two hydrophobic subsites contacting the side chains of the peptide Leu4 and Leu7 residues. An adjacent negatively charged subsite on the SH3 surface is likely to interact with the side chain of a basic residue at peptide position 10 that we show to be involved in binding. The peptide-binding site of the SH3 is characterized by large perturbations of amide chemical shifts when the peptide is added to the SH3 domain. The mGrb2 N-terminal SH3 domain structure in the complex is well-defined (backbone RMSD of 0.56 +/- 0.21 calculated over the backbone N, C alpha, and C atoms of residues 1-54). The structure of the peptide in the complex is less well-defined but displays a distinct orientation.(ABSTRACT TRUNCATED AT 250 WORDS)
- Yu H, Chen JK, Feng S, Dalgarno DC, Brauer AW, Schreiber SL
- Structural basis for the binding of proline-rich peptides to SH3 domains.
- Cell. 1994; 76: 933-45
- Display abstract
A common RXL motif was found in proline-rich ligands that were selected from a biased combinatorial peptide library on the basis of their ability to bind specifically to the SH3 domains from phosphatidylinositol 3-kinase (PI3K) or c-Src. The solution structure of the PI3K SH3 domain complexed to one of these ligands, RKLPPRPSK (RLP1), was determined. Structure-based mutations were introduced into the PI3K SH3 domain and the RLP1 ligand, and the influence of these mutations on binding was evaluated. We conclude that SH3 domains recognize proline-rich motifs possessing the left-handed type II polyproline (PPII) helix conformation. Two proline residues directly contact the receptor. Other prolines in the ligands appear to function as a molecular scaffold, promoting the formation of the PPII helix. Three nonproline residues consisting of combinations of arginine and leucine interact extensively with the SH3 domain and appear to confer ligand specificity.
- Booker GW et al.
- Solution structure and ligand-binding site of the SH3 domain of the p85 alpha subunit of phosphatidylinositol 3-kinase.
- Cell. 1993; 73: 813-22
- Display abstract
SH3 domains are found in proteins associated with receptor tyrosine kinase signal transduction complexes. The solution structure of the SH3 domain of the 85 kd regulatory subunit of phosphatidylinositol 3-kinase is shown to be a compact beta barrel consisting of five beta strands arranged in two beta sheets of three and two strands. The structure is similar to that of chicken brain alpha spectrin but represents a distinct class of SH3 domain, with an insertion between the second and third beta strands that may influence binding specificity. 1H chemical shift changes induced by complex formation with a synthetic peptide derived from the SH3-binding protein dynamin, together with amino acid sequence comparisons, suggest that the ligand-binding site consists of a hydrophobic surface flanked by two charged loops.
- Eck MJ, Shoelson SE, Harrison SC
- Recognition of a high-affinity phosphotyrosyl peptide by the Src homology-2 domain of p56lck.
- Nature. 1993; 362: 87-91
- Display abstract
The Src homology-2 (SH2) domains are modules of about 100 amino-acid residues that are found in many intracellular signal-transduction proteins. They bind phosphotyrosine-containing sequences with high affinity and specificity, recognizing phosphotyrosine in the context of the immediately adjacent polypeptide sequence. The protein p56lck (Lck) is a Src-like, lymphocyte-specific tyrosine kinase. A phosphopeptide library screen has recently been used to deduce an 'optimal' binding sequence for the Lck SH2 domain. There is selectivity for the residues Glu, Glu and Ile in the three positions C-terminal to the phosphotyrosine. An 11-residue phosphopeptide derived from the hamster polyoma middle-T antigen, EPQpYEEIPIYL, binds with an approximately 1 nM dissociation constant to the Lck SH2 (ref. 17), an affinity equivalent to that of the tightest known SH2-phosphopeptide complex. We report here the high-resolution crystallographic analysis of the Lck SH2 domain in complex with this phosphopeptide. Recent crystallographically derived structures of the Src SH2 domain in complex with low-affinity peptides, which do not contain the EEI consensus, and NMR-derived structures of unliganded Abl (ref. 19) and p85 (ref. 20) SH2 domains have revealed the conserved fold of the SH2 domain and the properties of a phosphotyrosine binding pocket. Our high-affinity complex shows the presence of a second pocket for the residue (pY + 3) three positions C-terminal to the phosphotyrosine (pY). The peptide is anchored by insertion of the pY and pY + 3 side chains into their pockets and by a network of hydrogen bonds to the peptide main chain. In the low-affinity phosphopeptide/Src complexes, the pY + 3 residues do not insert into the homologous binding pocket and the peptide main chain remains displaced from the surface of the domain.
- Kohda D et al.
- Solution structure of the SH3 domain of phospholipase C-gamma.
- Cell. 1993; 72: 953-60
- Display abstract
SH3 (Src homology 3) domains are found in many signaling proteins and appear to function as binding modules for cytoplasmic target proteins. The solution structure of the SH3 domain of human phospholipase C-gamma (PLC-gamma) was determined by two-dimensional 1H NMR analysis. This SH3 domain is composed of eight antiparallel beta strands consisting of two successive "Greek key" motifs, which form a barrel-like structure. The conserved aliphatic and aromatic residues form a hydrophobic pocket on the molecular surface, and the conserved carboxylic residues are localized to the periphery. The hydrophobic pocket may serve as a binding site for target proteins. Analysis of the slowly exchanging amide protons by NMR measurements indicates that despite containing a high content of beta structure, the SH3 domain of PLC-gamma is flexible.
- Koyama S, Yu H, Dalgarno DC, Shin TB, Zydowsky LD, Schreiber SL
- Structure of the PI3K SH3 domain and analysis of the SH3 family.
- Cell. 1993; 72: 945-52
- Display abstract
Src homology 3 (SH3) domains, which are found in many proteins involved in intracellular signal transduction, mediate specific protein-protein interactions. The three-dimensional structure of the SH3 domain in the p85 subunit of the phosphatidylinositol 3-kinase (PI3K) has been determined by multidimensional NMR methods. The molecule consists of four short helices, two beta turns, and two antiparallel beta sheets. The beta sheets are highly similar to corresponding regions in the SH3 domain of the tyrosine kinase Src, even though the sequence identity of the two domains is low. There is a unique 15 amino acid insert in PI3K that contains three short helices. There are substantial differences in the identity of the amino acids that make up the receptor site of SH3 domains. The results suggest that while the overall structures of the binding sites in the PI3K and Src SH3 domains are similar, their ligand binding properties may differ.
- Noble ME, Musacchio A, Saraste M, Courtneidge SA, Wierenga RK
- Crystal structure of the SH3 domain in human Fyn; comparison of the three-dimensional structures of SH3 domains in tyrosine kinases and spectrin.
- EMBO J. 1993; 12: 2617-24
- Display abstract
The Src-homology 3 (SH3) region is a protein domain consisting of approximately 60 residues. It occurs in a large number of eukaryotic proteins involved in signal transduction, cell polarization and membrane--cytoskeleton interactions. The function is unknown, but it is probably involved in specific protein--protein interactions. Here we report the crystal structure of the SH3 domain of Fyn (a Src family tyrosine kinase) at 1.9 A resolution. The crystals have two SH3 molecules per asymmetric unit. These two Fyn SH3 domains are not related by a local twofold axis. The crystal structures of spectrin and Fyn SH3 domains as well as the solution structure of the Src SH3 domain show that these all have the same basic fold. A protein domain which has the same topology as SH3 is present in the prokaryotic regulatory enzyme BirA. The comparison between the crystal structures of Fyn and spectrin SH3 domains shows that a conserved surface patch, consisting mainly of aromatic residues, is flanked by two hairpin-like loops (residues 94-104 and 114-118 in Fyn). These loops are different in tyrosine kinase and spectrin SH3 domains. They could modulate the binding properties of the aromatic surface.
- Ren R, Mayer BJ, Cicchetti P, Baltimore D
- Identification of a ten-amino acid proline-rich SH3 binding site.
- Science. 1993; 259: 1157-61
- Display abstract
The Src homology 3 (SH3) region is a small protein domain present in a very large group of proteins, including cytoskeletal elements and signaling proteins. It is believed that SH3 domains serve as modules that mediate protein-protein associations and, along with Src homology 2 (SH2) domains, regulate cytoplasmic signaling. The SH3 binding sites of two SH3 binding proteins were localized to a nine- or ten-amino acid stretch very rich in proline residues. Similar SH3 binding motifs exist in the formins, proteins that function in pattern formation in embryonic limbs of the mouse, and one subtype of the muscarinic acetylcholine receptor. Identification of the SH3 binding site provides a basis for understanding the interaction between the SH3 domains and their targets.
- Waksman G, Shoelson SE, Pant N, Cowburn D, Kuriyan J
- Binding of a high affinity phosphotyrosyl peptide to the Src SH2 domain: crystal structures of the complexed and peptide-free forms.
- Cell. 1993; 72: 779-90
- Display abstract
The crystal structure of the Src SH2 domain complexed with a high affinity 11-residue phosphopeptide has been determined at 2.7 A resolution by X-ray diffraction. The peptide binds in an extended conformation and makes primary interactions with the SH2 domain at six central residues: PQ(pY)EEI. The phosphotyrosine and the isoleucine are tightly bound by two well-defined pockets on the protein surface, resulting in a complex that resembles a two-pronged plug engaging a two-holed socket. The glutamate residues are in solvent-exposed environments in the vicinity of basic side chains of the SH2 domain, and the two N-terminal residues cap the phosphotyrosine-binding site. The crystal structure of Src SH2 in the absence of peptide has been determined at 2.5 A resolution, and comparison with the structure of the high affinity complex reveals only localized and relatively small changes.
- Weir HM, Kraulis PJ, Hill CS, Raine AR, Laue ED, Thomas JO
- Structure of the HMG box motif in the B-domain of HMG1.
- EMBO J. 1993; 12: 1311-9
- Display abstract
The conserved, abundant chromosomal protein HMG1 consists of two highly homologous, folded, basic DNA-binding domains, each of approximately 80 amino acid residues, and an acidic C-terminal tail. Each folded domain represents an 'HMG box', a sequence motif recently recognized in certain sequence-specific DNA-binding proteins and which also occurs in abundant HMG1-like proteins that bind to DNA without sequence specificity. The HMG box is defined by a set of highly conserved residues (most distinctively aromatic and basic) and appears to define a novel DNA-binding structural motif. We have expressed the HMG box region of the B-domain of rat HMG1 (residues 88-164 of the intact protein) in Escherichia coli and we describe here the determination of its structure by 2D 1H-NMR spectroscopy. There are three alpha-helices (residues 13-29, 34-48 and 50-74), which together account for approximately 75% of the total residues and contain many of the conserved basic and aromatic residues. Strikingly, the molecule is L-shaped, the angle of approximately 80 degrees between the two arms being defined by a cluster of conserved, predominantly aromatic, residues. The distinctive shape of the HMG box motif, which is distinct from hitherto characterized DNA-binding motifs, may be significant in relation to its recognition of four-way DNA junctions.
- Booker GW et al.
- Structure of an SH2 domain of the p85 alpha subunit of phosphatidylinositol-3-OH kinase.
- Nature. 1992; 358: 684-7
- Display abstract
Receptor protein-tyrosine kinases, through phosphorylation of specific tyrosine residues, generate high-affinity binding sites which direct assembly of multienzyme signalling complexes. Many of these signalling proteins, including phospholipase C gamma, GTPase-activating protein and phosphatidylinositol-3-OH kinase, contain src-homology 2 (SH2) domains, which bind with high affinity and specificity to tyrosine-phosphorylated sequences. The critical role played by SH2 domains in signalling has been highlighted by recent studies showing that mutation of specific phosphorylation sites on the platelet-derived growth factor receptor impair its association with phosphatidylinositol-3-OH kinase, preventing growth factor-induced mitogenesis. Here we report the solution structure of an isolated SH2 domain from the 85K regulatory subunit of phosphatidylinositol-3-OH kinase, determined using multidimensional nuclear magnetic resonance spectroscopy. The structure is characterized by a central region of beta-sheet flanked by two alpha-helices, with a highly flexible loop close to functionally important residues previously identified by site-directed mutagenesis.
- Musacchio A, Noble M, Pauptit R, Wierenga R, Saraste M
- Crystal structure of a Src-homology 3 (SH3) domain.
- Nature. 1992; 359: 851-5
- Display abstract
The Src-homologous SH3 domain is a small domain present in a large number of proteins that are involved in signal transduction, such as the Src protein tyrosine kinase, or in membrane-cytoskeleton interactions, but the function of SH3 is still unknown (reviewed in refs 1-3). Here we report the three-dimensional structure at 1.8 A resolution of the SH3 domain of the cytoskeletal protein spectrin expressed in Escherichia coli. The domain is a compact beta-barrel made of five antiparallel beta-strands. The amino acids that are conserved in the SH3 sequences are located close to each other on one side of the molecule. This surface is rich in aromatic and carboxylic amino acids, and is distal to the region of the molecule where the N and C termini reside and where SH3 inserts into the alpha-spectrin chain. We suggest that a protein ligand binds to this conserved surface of SH3.
- Waksman G et al.
- Crystal structure of the phosphotyrosine recognition domain SH2 of v-src complexed with tyrosine-phosphorylated peptides.
- Nature. 1992; 358: 646-53
- Display abstract
Three-dimensional structures of complexes of the SH2 domain of the v-src oncogene product with two phosphotyrosyl peptides have been determined by X-ray crystallography at resolutions of 1.5 and 2.0 A, respectively. A central antiparallel beta-sheet in the structure is flanked by two alpha-helices, with peptide binding mediated by the sheet, intervening loops and one of the helices. The specific recognition of phosphotyrosine involves amino-aromatic interactions between lysine and arginine side chains and the ring system in addition to hydrogen-bonding interactions with the phosphate.
- Yu H, Rosen MK, Shin TB, Seidel-Dugan C, Brugge JS, Schreiber SL
- Solution structure of the SH3 domain of Src and identification of its ligand-binding site.
- Science. 1992; 258: 1665-8
- Display abstract
The Src homology 3 (SH3) region is a protein domain of 55 to 75 amino acids found in many cytoplasmic proteins, including those that participate in signal transduction pathways. The solution structure of the SH3 domain of the tyrosine kinase Src was determined by multidimensional nuclear magnetic resonance methods. The molecule is composed of two short three-stranded anti-parallel beta sheets packed together at approximately right angles. Studies of the SH3 domain bound to proline-rich peptide ligands revealed a hydrophobic binding site on the surface of the protein that is lined with the side chains of conserved aromatic amino acids.
- Otsu M et al.
- Characterization of two 85 kd proteins that associate with receptor tyrosine kinases, middle-T/pp60c-src complexes, and PI3-kinase.
- Cell. 1991; 65: 91-104
- Display abstract
Affinity-purified bovine brain phosphatidylinositol 3-kinase (PI3-kinase) contains two major proteins of 85 and 110 kd. Amino acid sequence analysis and cDNA cloning reveals two related 85 kd proteins (p85 alpha and p85 beta), which both contain one SH3 and two SH2 regions (src homology regions). When expressed, these 85 kd proteins bind to and are substrates for tyrosine-phosphorylated receptor kinases and the polyoma virus middle-T antigen/pp60c-src complex, but lack PI3-kinase activity. However, an antiserum raised against p85 beta immunoprecipitates PI3-kinase activity. The active PI3-kinase complex containing p85 alpha or p85 beta and the 110 kd protein binds to PDGF but not EGF receptors. p85 alpha and p85 beta may mediate specific PI3-kinase interactions with a subset of tyrosine kinases.
- Disease (disease genes where sequence variants are found in this domain)
-
SwissProt sequences and OMIM curated human diseases associated with missense mutations within the SH3 domain.
Protein Disease Unconventional myosin-VIIa (Q13402) (SMART) OMIM:276903: Usher syndrome, type 1B ; Deafness, autosomal recessive 2, neurosensory
OMIM:600060: Deafness, autosomal dominant 11, neurosensory
OMIM:601317:Peroxisomal membrane protein PEX13 (Q92968) (SMART) OMIM:601789: Zellweger syndrome
OMIM:214100: Adrenoleukodystrophy, neonatal
OMIM:202370: - Metabolism (metabolic pathways involving proteins which contain this domain)
-
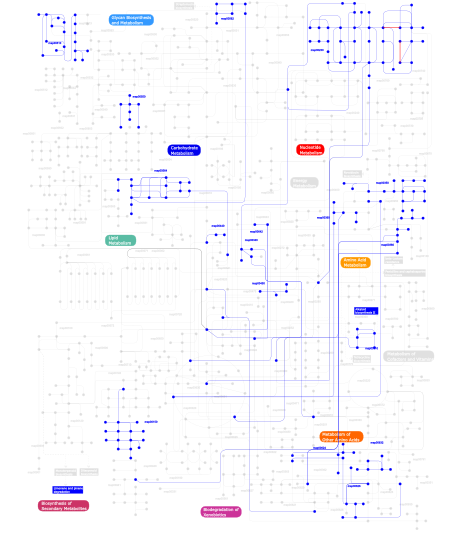
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 5.36 map04012 ErbB signaling pathway 5.14 map04660 T cell receptor signaling pathway 5.08 map04670 Leukocyte transendothelial migration 5.02 map04010 MAPK signaling pathway 4.85 map04810 Regulation of actin cytoskeleton 4.63 map04664 Fc epsilon RI signaling pathway 4.51 map04510 Focal adhesion 4.12 map04530 Tight junction 3.95 map04650 Natural killer cell mediated cytotoxicity 3.72 map04360 Axon guidance 2.82 map05120 Epithelial cell signaling in Helicobacter pylori infection 2.82 map04910 Insulin signaling pathway 2.77 map04662 B cell receptor signaling pathway 2.71 map05220 Chronic myeloid leukemia 2.60 map04520 Adherens junction 2.54 map04630 Jak-STAT signaling pathway 2.26 map05214 Glioma 2.26 map05223 Non-small cell lung cancer 2.26 map05211 Renal cell carcinoma 2.20 map04370 VEGF signaling pathway 1.75 map04070 Phosphatidylinositol signaling system 1.47 map04540 Gap junction 1.41 map05213 Endometrial cancer 1.41 map05221 Acute myeloid leukemia 1.41 map05210 Colorectal cancer 1.41 map05215 Prostate cancer 1.30 map05212 Pancreatic cancer 1.07 map04912 GnRH signaling pathway 1.07 map05040 Huntington's disease 0.90  map00562
map00562Inositol phosphate metabolism 0.85 map04210 Apoptosis 0.85 map04930 Type II diabetes mellitus 0.85 map04914 Progesterone-mediated oocyte maturation 0.85 map05222 Small cell lung cancer 0.85 map04020 Calcium signaling pathway 0.85 map04620 Toll-like receptor signaling pathway 0.85 map04150 mTOR signaling pathway 0.85 map05218 Melanoma 0.56 map01030 Glycan structures - biosynthesis 1 0.56 map04320 Dorso-ventral axis formation 0.56 map04730 Long-term depression 0.56  map00510
map00510N-Glycan biosynthesis 0.56 map00533 Keratan sulfate biosynthesis 0.45 map04110 Cell cycle 0.40  map00350
map00350Tyrosine metabolism 0.40  map00340
map00340Histidine metabolism 0.40 map05050 Dentatorubropallidoluysian atrophy (DRPLA) 0.34  map00230
map00230Purine metabolism 0.28  map00642
map00642Ethylbenzene degradation 0.28  map00632
map00632Benzoate degradation via CoA ligation 0.28  map00564
map00564Glycerophospholipid metabolism 0.28  map00360
map00360Phenylalanine metabolism 0.28 map00903 Limonene and pinene degradation 0.28 map00960 Alkaloid biosynthesis II 0.28  map00310
map00310Lysine degradation 0.28 map03320 PPAR signaling pathway 0.28  map00624
map006241- and 2-Methylnaphthalene degradation 0.28  map00550
map00550Peptidoglycan biosynthesis 0.11  map00150
map00150Androgen and estrogen metabolism 0.11  map00626
map00626Naphthalene and anthracene degradation 0.11  map00380
map00380Tryptophan metabolism 0.11  map00440
map00440Aminophosphonate metabolism 0.11  map00450
map00450Selenoamino acid metabolism 0.06 map04610 Complement and coagulation cascades 0.06 map00511 N-Glycan degradation 0.06 map02010 ABC transporters - General 0.06 map01032 Glycan structures - degradation This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with SH3 domain which could be assigned to a KEGG orthologous group, and not all proteins containing SH3 domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of SH3 domains in PDB
PDB code Main view Title 1a0n 
NMR STUDY OF THE SH3 DOMAIN FROM FYN PROTO-ONCOGENE TYROSINE KINASE COMPLEXED WITH THE SYNTHETIC PEPTIDE P2L CORRESPONDING TO RESIDUES 91-104 OF THE P85 SUBUNIT OF PI3-KINASE, FAMILY OF 25 STRUCTURES 1abo 
CRYSTAL STRUCTURE OF THE COMPLEX OF THE ABL TYROSINE KINASE SH3 DOMAIN WITH 3BP-1 SYNTHETIC PEPTIDE 1abq 
CRYSTAL STRUCTURE OF THE UNLIGANDED ABL TYROSINE KINASE SH3 DOMAIN 1ad5 
SRC FAMILY KINASE HCK-AMP-PNP COMPLEX 1aey 
ALPHA-SPECTRIN SRC HOMOLOGY 3 DOMAIN, SOLUTION NMR, 15 STRUCTURES 1aoj 
THE SH3 DOMAIN OF EPS8 EXISTS AS A NOVEL INTERTWINED DIMER 1ark 
SH3 DOMAIN FROM HUMAN NEBULIN, NMR, 15 STRUCTURES 1avz 
V-1 NEF PROTEIN IN COMPLEX WITH WILD TYPE FYN SH3 DOMAIN 1awj 
INTRAMOLECULAR ITK-PROLINE COMPLEX, NMR, MINIMIZED AVERAGE STRUCTURE 1awo 
THE SOLUTION NMR STRUCTURE OF ABL SH3 AND ITS RELATIONSHIP TO SH2 IN THE SH(32) CONSTRUCT, 20 STRUCTURES 1aww 
SH3 DOMAIN FROM BRUTON'S TYROSINE KINASE, NMR, 42 STRUCTURES 1awx 
SH3 DOMAIN FROM BRUTON'S TYROSINE KINASE, NMR, MINIMIZED AVERAGE STRUCTURE 1aze 
NMR STRUCTURE OF THE COMPLEX BETWEEN THE C32S-Y7V MUTANT OF THE NSH3 DOMAIN OF GRB2 WITH A PEPTIDE FROM SOS, 10 STRUCTURES 1azg 
NMR STUDY OF THE SH3 DOMAIN FROM FYN PROTO-ONCOGENE TYROSINE KINASE KINASE COMPLEXED WITH THE SYNTHETIC PEPTIDE P2L CORRESPONDING TO RESIDUES 91-104 OF THE P85 SUBUNIT OF PI3-KINASE, MINIMIZED AVERAGE (PROBMAP) STRUCTURE 1b07 
CRK SH3 DOMAIN COMPLEXED WITH PEPTOID INHIBITOR 1bb9 
CRYSTAL STRUCTURE OF THE SH3 DOMAIN FROM RAT AMPHIPHYSIN 2 1bbz 
CRYSTAL STRUCTURE OF THE ABL-SH3 DOMAIN COMPLEXED WITH A DESIGNED HIGH-AFFINITY PEPTIDE LIGAND: IMPLICATIONS FOR SH3-LIGAND INTERACTIONS 1bk2 
A-SPECTRIN SH3 DOMAIN D48G MUTANT 1bu1 
SRC FAMILY KINASE HCK SH3 DOMAIN 1cka 
STRUCTURAL BASIS FOR THE SPECIFIC INTERACTION OF LYSINE-CONTAINING PROLINE-RICH PEPTIDES WITH THE N-TERMINAL SH3 DOMAIN OF C-CRK 1ckb 
STRUCTURAL BASIS FOR THE SPECIFIC INTERACTION OF LYSINE-CONTAINING PROLINE-RICH PEPTIDES WITH THE N-TERMINAL SH3 DOMAIN OF C-CRK 1csk 
THE CRYSTAL STRUCTURE OF HUMAN CSKSH3: STRUCTURAL DIVERSITY NEAR THE RT-SRC AND N-SRC LOOP 1e6g 
A-SPECTRIN SH3 DOMAIN D48G MUTANT 1e6h 
A-SPECTRIN SH3 DOMAIN A11V, M25I, V44I, V58L MUTANTS 1e7o 
A-SPECTRIN SH3 DOMAIN A11V, V23L, M25V, V44I, V58L MUTATIONS 1efn 
HIV-1 NEF PROTEIN IN COMPLEX WITH R96I MUTANT FYN SH3 DOMAIN 1fmk 
CRYSTAL STRUCTURE OF HUMAN TYROSINE-PROTEIN KINASE C-SRC 1fyn 
PHOSPHOTRANSFERASE 1g2b 
ALPHA-SPECTRIN SRC HOMOLOGY 3 DOMAIN, CIRCULAR PERMUTANT, CUT AT N47-D48 1g83 
CRYSTAL STRUCTURE OF FYN SH3-SH2 1gbq 
SOLUTION NMR STRUCTURE OF THE GRB2 N-TERMINAL SH3 DOMAIN COMPLEXED WITH A TEN-RESIDUE PEPTIDE DERIVED FROM SOS DIRECT REFINEMENT AGAINST NOES, J-COUPLINGS, AND 1H AND 13C CHEMICAL SHIFTS, MINIMIZED AVERAGE STRUCTURE 1gbr 
ORIENTATION OF PEPTIDE FRAGMENTS FROM SOS PROTEINS BOUND TO THE N-TERMINAL SH3 DOMAIN OF GRB2 DETERMINED BY NMR SPECTROSCOPY 1gcp 
CRYSTAL STRUCTURE OF VAV SH3 DOMAIN 1gcq 
CRYSTAL STRUCTURE OF VAV AND GRB2 SH3 DOMAINS 1gfc 
SOLUTION STRUCTURE AND LIGAND-BINDING SITE OF THE C-TERMINAL SH3 DOMAIN OF GRB2 1gfd 
SOLUTION STRUCTURE AND LIGAND-BINDING SITE OF THE C-TERMINAL SH3 DOMAIN OF GRB2 1gl5 
NMR structure of the SH3 domain from the Tec protein tyrosine kinase 1gri 
GRB2 1h3h 
Structural Basis for Specific Recognition of an RxxK-containing SLP-76 peptide by the Gads C-terminal SH3 domain 1h8k 
A-SPECTRIN SH3 DOMAIN A11V, V23L, M25V, V53I, V58L MUTANT 1h92 
SH3 domain of human Lck tyrosine kinase 1hd3 
A-SPECTRIN SH3 DOMAIN F52Y MUTANT 1hjd 
Melanoma inhibitory activity (MIA) protein 1hsq 
SOLUTION STRUCTURE OF THE SH3 DOMAIN OF PHOSPHOLIPASE CGAMMA 1i07 
EPS8 SH3 DOMAIN INTERTWINED DIMER 1i0c 
EPS8 SH3 CLOSED MONOMER 1i1j 
STRUCTURE OF MELANOMA INHIBITORY ACTIVITY PROTEIN: A MEMBER OF A NEW FAMILY OF SECRETED PROTEINS 1io6 
GROWTH FACTOR RECEPTOR-BOUND PROTEIN 2 (GRB2) C-TERMINAL SH3 DOMAIN COMPLEXED WITH A LIGAND PEPTIDE (NMR, MINIMIZED MEAN STRUCTURE) 1j3t 
Solution structure of the second SH3 domain of human intersectin 2 (KIAA1256) 1jeg 
Solution structure of the SH3 domain from C-terminal Src Kinase complexed with a peptide from the tyrosine phosphatase PEP 1jo8 
Structural analysis of the yeast actin binding protein Abp1 SH3 domain 1jqq 
Crystal structure of Pex13p(301-386) SH3 domain 1ju5 
Ternary complex of an Crk SH2 domain, Crk-derived phophopeptide, and Abl SH3 domain by NMR spectroscopy 1jxm 
CRYSTAL STRUCTURE OF THE GMP BOUND SH3-HOOK-GK FRAGMENT OF PSD-95 1jxo 
Crystal Structure of the SH3-HOOK-GK Fragment of PSD-95 1k0x 
Solution Structure of Melanoma Inhibitory Activity Protein 1k1z 
Solution structure of N-terminal SH3 domain mutant(P33G) of murine Vav 1k4u 
Solution structure of the C-terminal SH3 domain of p67phox complexed with the C-terminal tail region of p47phox 1k76 
Solution Structure of the C-terminal Sem-5 SH3 Domain (Minimized Average Structure) 1k9a 
Crystal structure analysis of full-length carboxyl-terminal Src kinase at 2.5 A resolution 1kfz 
Solution Structure of C-terminal Sem-5 SH3 Domain (Ensemble of 16 Structures) 1kik 
SH3 Domain of Lymphocyte Specific Kinase (LCK) 1kjw 
SH3-Guanylate Kinase Module from PSD-95 1ksw 
Structure of Human c-Src Tyrosine Kinase (Thr338Gly Mutant) in Complex with N6-benzyl ADP 1lck 
SH3-SH2 DOMAIN FRAGMENT OF HUMAN P56-LCK TYROSINE KINASE COMPLEXED WITH THE 10 RESIDUE SYNTHETIC PHOSPHOTYROSYL PEPTIDE TEGQPYQPQPA 1m27 
Crystal structure of SAP/FynSH3/SLAM ternary complex 1m30 
Solution structure of N-terminal SH3 domain from oncogene protein c-Crk 1m3a 
Solution structure of a circular form of the truncated N-terminal SH3 domain from oncogene protein c-Crk. 1m3b 
Solution structure of a circular form of the N-terminal SH3 domain (A134C, E135G, R191G mutant) from oncogene protein c-Crk. 1m3c 
Solution structure of a circular form of the N-terminal SH3 domain (E132C, E133G, R191G mutant) from oncogene protein c-Crk 1m8m 
SOLID-STATE MAS NMR STRUCTURE OF THE A-SPECTRIN SH3 DOMAIN 1muz 
NMR STRUCTURE OF THE TUMOR SUPPRESSOR BIN1: ALTERNATIVE SPLICING IN MELANOMA AND INTERACTION WITH C-MYC 1mv0 
NMR STRUCTURE OF THE TUMOR SUPPRESSOR BIN1: ALTERNATIVE SPLICING IN MELANOMA AND INTERACTION WITH C-MYC 1mv3 
NMR STRUCTURE OF THE TUMOR SUPPRESSOR BIN1: ALTERNATIVE SPLICING IN MELANOMA AND INTERACTION WITH C-MYC 1n5z 
Complex structure of Pex13p SH3 domain with a peptide of Pex14p 1neb 
SH3 DOMAIN FROM HUMAN NEBULIN, NMR, MINIMIZED AVERAGE STRUCTURE 1neg 
Crystal Structure Analysis of N-and C-terminal labeled SH3-domain of alpha-Chicken Spectrin 1ng2 
Structure of autoinhibited p47phox 1nlo 
STRUCTURE OF SIGNAL TRANSDUCTION PROTEIN, NMR, MINIMIZED AVERAGE STRUCTURE 1nlp 
STRUCTURE OF SIGNAL TRANSDUCTION PROTEIN, NMR, MINIMIZED AVERAGE STRUCTURE 1nm7 
Solution structure of the ScPex13p SH3 domain 1nyf 
NMR STUDY OF THE SH3 DOMAIN FROM FYN PROTO-ONCOGENE TYROSINE KINASE, MINIMIZED AVERAGE (PROBMAP) STRUCTURE 1nyg 
NMR STUDY OF THE SH3 DOMAIN FROM FYN PROTO-ONCOGENE TYROSINE KINASE, FAMILY OF 20 STRUCTURES 1oeb 
Mona/Gads SH3C domain 1oot 
Crystal structure of the SH3 domain from a S. cerevisiae hypothetical 40.4 kDa protein at 1.39 A resolution 1opk 
Structural basis for the auto-inhibition of c-Abl tyrosine kinase 1opl 
Structural basis for the auto-inhibition of c-Abl tyrosine kinase 1ov3 
Structure of the p22phox-p47phox complex 1pht 
PHOSPHATIDYLINOSITOL 3-KINASE P85-ALPHA SUBUNIT SH3 DOMAIN, RESIDUES 1-85 1pks 
STRUCTURE OF THE PI3K SH3 DOMAIN AND ANALYSIS OF THE SH3 FAMILY 1pkt 
STRUCTURE OF THE PI3K SH3 DOMAIN AND ANALYSIS OF THE SH3 FAMILY 1pnj 
SOLUTION STRUCTURE AND LIGAND-BINDING SITE OF THE SH3 DOMAIN OF THE P85ALPHA SUBUNIT OF PHOSPHATIDYLINOSITOL 3-KINASE 1prl 
TWO BINDING ORIENTATIONS FOR PEPTIDES TO SRC SH3 DOMAIN: DEVELOPMENT OF A GENERAL MODEL FOR SH3-LIGAND INTERACTIONS 1prm 
TWO BINDING ORIENTATIONS FOR PEPTIDES TO SRC SH3 DOMAIN: DEVELOPMENT OF A GENERAL MODEL FOR SH3-LIGAND INTERACTIONS 1pwt 
THERMODYNAMIC ANALYSIS OF ALPHA-SPECTRIN SH3 AND TWO OF ITS CIRCULAR PERMUTANTS WITH DIFFERENT LOOP LENGTHS: DISCERNING THE REASONS FOR RAPID FOLDING IN PROTEINS 1qcf 
CRYSTAL STRUCTURE OF HCK IN COMPLEX WITH A SRC FAMILY-SELECTIVE TYROSINE KINASE INHIBITOR 1qkw 
Alpha-spectrin Src Homology 3 domain, N47G mutant in the distal loop. 1qkx 
Alpha-spectrin Src Homology 3 domain, N47A mutant in the distal loop. 1qly 
NMR Study of the SH3 Domain From Bruton's Tyrosine Kinase, 20 Structure 1qwe 
C-SRC SH3 DOMAIN COMPLEXED WITH LIGAND APP12 1qwf 
C-SRC SH3 DOMAIN COMPLEXED WITH LIGAND VSL12 1ri9 
Structure of a helically extended SH3 domain of the T cell adapter protein ADAP 1rlp 
TWO BINDING ORIENTATIONS FOR PEPTIDES TO SRC SH3 DOMAIN: DEVELOPMENT OF A GENERAL MODEL FOR SH3-LIGAND INTERACTIONS 1rlq 
TWO BINDING ORIENTATIONS FOR PEPTIDES TO SRC SH3 DOMAIN: DEVELOPMENT OF A GENERAL MODEL FOR SH3-LIGAND INTERACTIONS 1ruw 
Crystal structure of the SH3 domain from S. cerevisiae Myo3 1s1n 
SH3 domain of human nephrocystin 1sem 
STRUCTURAL DETERMINANTS OF PEPTIDE-BINDING ORIENTATION AND OF SEQUENCE SPECIFICITY IN SH3 DOMAINS 1shf 
CRYSTAL STRUCTURE OF THE SH3 DOMAIN IN HUMAN FYN; COMPARISON OF THE THREE-DIMENSIONAL STRUCTURES OF SH3 DOMAINS IN TYROSINE KINASES AND SPECTRIN 1shg 
CRYSTAL STRUCTURE OF A SRC-HOMOLOGY 3 (SH3) DOMAIN 1spk 
Solution Structure of RSGI RUH-010, an SH3 Domain from Mouse cDNA 1srl 
1H AND 15N ASSIGNMENTS AND SECONDARY STRUCTURE OF THE SRC SH3 DOMAIN 1srm 
1H AND 15N ASSIGNMENTS AND SECONDARY STRUCTURE OF THE SRC SH3 DOMAIN 1ssh 
Crystal structure of the SH3 domain from a S. cerevisiae hypothetical 40.4 kDa protein in complex with a peptide 1t0h 
Crystal structure of the Rattus norvegicus voltage gated calcium channel beta subunit isoform 2a 1t0j 
Crystal structure of a complex between voltage-gated calcium channel beta2a subunit and a peptide of the alpha1c subunit 1t3l 
Structural Analysis of the Voltage-Dependent Calcium Channel Beta Subunit Functional Core in Complex with Alpha1 Interaction Domain 1t3s 
Structural Analysis of the Voltage-Dependent Calcium Channel Beta Subunit Functional Core 1tg0 
0.97-A structure of the SH3 domain of bbc1 1tuc 
ALPHA-SPECTRIN SRC HOMOLOGY 3 DOMAIN, CIRCULAR PERMUTANT, CUT AT S19-P20 1tud 
ALPHA-SPECTRIN SRC HOMOLOGY 3 DOMAIN, CIRCULAR PERMUTANT, CUT AT N47-D48 1u06 
crystal structure of chicken alpha-spectrin SH3 domain 1u3o 
Solution structure of rat Kalirin N-terminal SH3 domain 1u5s 
NMR structure of the complex between Nck-2 SH3 domain and PINCH-1 LIM4 domain 1udl 
The solution structure of the fifth SH3 domain of intersectin 2 (KIAA1256) 1ue9 
Solution structure of the fourth SH3 domain of human intersectin 2 (KIAA1256) 1uec 
Crystal structure of autoinhibited form of tandem SH3 domain of p47phox 1uff 
Solution structure of the first SH3 domain of human intersectin2 (KIAA1256) 1ug1 
SH3 domain of Hypothetical protein BAA76854.1 1ugv 
Solution structure of the SH3 domain of human olygophrein-1 like protein (KIAA0621) 1uhc 
Solution Structure of RSGI RUH-002, a SH3 Domain of KIAA1010 protein [Homo sapiens] 1uhf 
Solution Structure of the third SH3 domain of human intersectin 2(KIAA1256) 1uj0 
Crystal Structure of STAM2 SH3 domain in complex with a UBPY-derived peptide 1ujy 
Solution structure of SH3 domain in Rac/Cdc42 guanine nucleotide exchange factor(GEF) 6 1uti 
Mona/Gads SH3C in complex with HPK derived peptide 1uue 
a-SPECTRIN SH3 DOMAIN (V44T, D48G MUTANT) 1v1c 
Solution Structure of the SH3 domain of Obscurin 1va7 
Yeast Myo3 SH3 domain, triclinic crystal form 1vyt 
beta3 subunit complexed with aid 1vyu 
Beta3 subunit of Voltage-gated Ca2+-channel 1vyv 
beta4 subunit of Ca2+ channel 1w1f 
SH3 DOMAIN OF HUMAN LYN TYROSINE KINASE 1w6x 
SH3 domain of p40phox, component of the NADPH oxidase 1w70 
SH3 domain of p40phox complexed with C-terminal polyProline of p47phox, components of the NADPH oxidase 1wa7 
SH3 DOMAIN OF HUMAN LYN TYROSINE KINASE IN COMPLEX WITH A HERPESVIRAL LIGAND 1wdx 
Yeast BBC1 SH3 domain, triclinic crystal form 1wfw 
Solution structure of SH3 domain of mouse Kalirin-9a protein 1wi7 
Solution structure of the SH3 domain of SH3-domain kinase binding protein 1 1wie 
Solution structure of the first SH3 domain of KIAA0318 protein 1wlp 
Solution Structure Of The P22Phox-P47Phox Complex 1wx6 
Solution structure of the SH3 domain of the human cytoplasmic protein NCK2 1wxb 
Solution structure of the SH3 domain from human epidermal growth factor receptor pathway substrate 8-like protein 1wxt 
Solution structure of the SH3 domain of human hypothetical protein FLJ21522 1wxu 
Solution structure of the SH3 domain of mouse peroxisomal biogenesis factor 13 1wyx 
The Crystal Structure of the p130Cas SH3 Domain at 1.1 A Resolution 1x27 
Crystal Structure of Lck SH2-SH3 with SH2 binding site of p130Cas 1x2k 
Solution Structure of the SH3 Domain of Human osteoclast stimulating factor 1 (OSTF1) 1x2p 
Solution structure of the SH3 domain of the Protein arginine N-methyltransferase 2 1x2q 
Solution structure of the SH3 domain of the Signal transducing adaptor molecule 2 1x43 
Solution structure of the SH3 domain of Endophilin B1 (Sh3g1b1) 1x69 
Solution structures of the SH3 domain of human Src substrate cortactin 1x6b 
Solution structures of the SH3 domain of human rho guanine exchange factor (GEF) 16 1x6g 
Solution structures of the SH3 domain of human megakaryocyte-associated tyrosine-protein kinase. 1y0m 
Crystal structure of of the SH3 domain of phospholipase C Gamma-1 1y57 
Structure of unphosphorylated c-Src in complex with an inhibitor 1ycs 
P53-53BP2 COMPLEX 1yn8 
SH3 domain of yeast NBP2 1ynz 
SH3 domain of yeast Pin3 1yp5 
Yeast Myo5 SH3 domain, tetragonal crystal form 1ywo 
Phospholipase Cgamma1 SH3 in complex with a SLP-76 motif 1ywp 
Phospholipase Cgamma1 SH3 1z9q 
Solution structure of SH3 domain of p40phox 1z9z 
Crystal structure of yeast sla1 SH3 domain 3 1zbj 
Inferential Structure Determination of the Fyn SH3 domain using NOESY data from a 15N,H2 enriched protein 1zlm 
Crystal structure of the SH3 domain of human osteoclast stimulating factor 1zsg 
beta PIX-SH3 complexed with an atypical peptide from alpha-PAK 1zuk 
Yeast BBC1 Sh3 domain complexed with a peptide from Las17 1zuu 
Crystal structure of the yeast Bzz1 first SH3 domain at 0.97-A resolution 1zuy 
High-resolution structure of yeast Myo5 SH3 domain 1zx6 
High-resolution crystal structure of yeast Pin3 SH3 domain 2a08 
Structure of the yeast YHH6 SH3 domain 2a28 
Atomic-resolution crystal structure of the second SH3 domain of yeast Bzz1 determined from a pseudomerohedrally twinned crystal 2a36 
Solution structure of the N-terminal SH3 domain of DRK 2a37 
Solution structure of the T22G mutant of N-terminal SH3 domain of DRK (DRKN SH3 DOMAIN) 2abl 
SH3-SH2 DOMAIN FRAGMENT OF HUMAN BCR-ABL TYROSINE KINASE 2ak5 
beta PIX-SH3 complexed with a Cbl-b peptide 2azs 
NMR structure of the N-terminal SH3 domain of Drk (calculated without NOE restraints) 2azv 
Solution structure of the T22G mutant of N-terminal SH3 domain of DRK (calculated without NOEs) 2b86 
Solution structure of the first Src homology 3 domain of Nck2 2btt 
NMR Structure of MYO3-SH3 domain from Myosin-typeI from S. cerevisiae 2bz8 
N-terminal Sh3 domain of CIN85 bound to Cbl-b peptide 2bzx 
Atomic model of CrkL-SH3C monomer 2bzy 
Homodimer of CrkL-SH3C domain 2c0i 
Src family kinase Hck with bound inhibitor A-420983 2c0o 
Src family kinase Hck with bound inhibitor A-770041 2c0t 
Src family kinase Hck with bound inhibitor A-641359 2cdt 
alpha-SPECTRIN SH3 DOMAIN A56S MUTANT 2cre 
Solution structure of RSGI RUH-036, an SH3 domain from human cDNA 2csi 
Solution structure of the third SH3 domain of human RIM-binding protein 2 2csq 
Solution structure of the second SH3 domain of human RIM-binding protein 2 2ct3 
Solution Structure of the SH3 domain of the Vinexin protein 2ct4 
Solution Strutcure of the SH3 domain of the Cdc42-interacting protein 4 2cub 
Solution structure of the SH3 domain of the human cytoplasmic protein Nck1 2cuc 
Solution structure of the SH3 domain of the mouse hypothetical protein SH3RF2 2cud 
Solution structure of the SH3 domain of the human SRC-like adopter protein (SLAP) 2d0n 
Crystal structure of the C-terminal SH3 domain of the adaptor protein GADS in complex with SLP-76 motif peptide reveals a unique SH3-SH3 interaction 2d1x 
The crystal structure of the cortactin-SH3 domain and AMAP1-peptide complex 2d8h 
Solution structure of the SH3 domain of Hypothetical protein SH3YL1 2d8j 
Solution structure of the SH3 domain of Fyn-related kinase 2da9 
Solution structure of the third SH3 domain of SH3-domain kinase binding protein 1 (Regulator of ubiquitous kinase, Ruk) 2dbk 
Solution structures of the SH3 domain of human Crk-like protein 2dbm 
Solution structures of the SH3 domain of human SH3-containing GRB2-like protein 2 2de0 
Crystal structure of human alpha 1,6-fucosyltransferase, FUT8 2df6 
Crystal Structure of the SH3 Domain of betaPIX in Complex with a High Affinity Peptide from PAK2 2dil 
Solution structure of the SH3 domain of the human Proline-serine-threonine phosphatase-interacting protein 1 2djq 
The solution structure of the first SH3 domain of mouse SH3 domain containing ring finger 2 2dl3 
Solution structure of the first SH3 domain of human sorbin and Sh3 domain-containing protein 1 2dl4 
Solution structure of the first SH3 domain of Stac protein 2dl5 
Solution structure of the first SH3 domain of human KIAA0769 protein 2dl7 
Solution structure of the second SH3 domain of human KIAA0769 protein 2dl8 
Solution structure of the SH3 domain of human SLIT-ROBO Rho GTPase-activating protein 2 2dlm 
Solution structure of the first SH3 domain of human vinexin 2dlp 
Solution structure of the SH3 domain of human KIAA1783 protein 2dm1 
Solution structure of the second SH3 domain of human protein vav-2 2dmo 
Refined solution structure of the 1st SH3 domain from human Neutrophil cytosol factor 2 (NCF-2) 2dnu 
Solution structure of RSGI RUH-061, a SH3 domain from human 2drk 
Acanthamoeba myosin I SH3 domain bound to Acan125 2drm 
Acanthamoeba myosin I SH3 domain bound to Acan125 2dvj 
phosphorylated Crk-II 2dx1 
Crystal structure of RhoGEF protein Asef 2dyb 
The crystal structure of human p40(phox) 2e5k 
Solution structure of SH3 domain in Suppressor of T-cell receptor signaling 1 2ebp 
Solution structure of the SH3 domain from human SAM and SH3 domain containing protein 1 2ecz 
Solution structure of the SH3 domain of Sorbin and SH3 domain-containing protein 1 2ed0 
Solution structure of the SH3 domain of Abl interactor 2 (Abelson interactor 2) 2ed1 
Solution structure of the SH3 domain of 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase-activating protein 2ega 
Solution structure of the first SH3 domain from human KIAA0418 protein 2egc 
Solution structure of the fifth SH3 domain from human KIAA0418 protein 2ege 
Solution structure of the third SH3 domain from human KIAA1666 protein 2ekh 
Solution structures of the SH3 domain of human KIAA0418 2enm 
Solution structure of the SH3 domain from mouse sorting nexin-9 2epd 
Solution structure of SH3 domain in Rho-GTPase-activating protein 4 2eqi 
Solution structure of the SH3 domain from Phospholipase C, gamma 2 2esw 
Atomic structure of the N-terminal SH3 domain of mouse beta PIX,p21-activated kinase (PAK)-interacting exchange factor 2ew3 
Solution Structure Of The SH3 Domain Of Human SH3GL3 2eyw 
N-terminal SH3 domain of CT10-Regulated Kinase 2eyx 
C-Terminal SH3 domain of CT10-Regulated Kinase 2eyy 
CT10-Regulated Kinase isoform I 2eyz 
CT10-Regulated Kinase isoform II 2f2v 
alpha-spectrin SH3 domain A56G mutant 2f2w 
alpha-spectrin SH3 domain R21A mutant 2f2x 
alpha-spectrin SH3 domain R21G mutant 2fei 
Solution structure of the second SH3 domain of Human CMS protein 2fo0 
Organization of the SH3-SH2 Unit in Active and Inactive Forms of the c-Abl Tyrosine Kinase 2fpd 
Sad structure determination: crystal structure of the intrinsic dimerization sh3 domain of the ib1 scaffold protein 2fpe 
Conserved dimerization of the ib1 src-homology 3 domain 2fpf 
Crystal structure of the ib1 sh3 dimer at low resolution 2frw 
Solution structure of the second SH3 domain of human adaptor protein NCK2 2fry 
Solution structure of the third SH3 domain of human NCK2 adaptor protein 2g6f 
Crystal Structure of the SH3 Domain of betaPIX in Complex with a High Affinity Peptide from PAK2 2gbq 
SOLUTION NMR STRUCTURE OF THE GRB2 N-TERMINAL SH3 DOMAIN COMPLEXED WITH A TEN-RESIDUE PEPTIDE DERIVED FROM SOS DIRECT REFINEMENT AGAINST NOES, J-COUPLINGS, AND 1H AND 13C CHEMICAL SHIFTS, 15 STRUCTURES 2ggr 
Solution structure of the C-terminal SH3 domain of c-CrkII 2gnc 
Crystal structure of srGAP1 SH3 domain in the slit-robo signaling pathway 2gqi 
Solution structure of the SH3 domain of human Ras GTPase-activating protein 1 2gtj 
Reduced form of ADAP hSH3-N-domain 2gto 
Oxidized form of ADAP hSH3-N 2h8h 
Src kinase in complex with a quinazoline inhibitor 2hck 
SRC FAMILY KINASE HCK-QUERCETIN COMPLEX 2hda 
Yes SH3 domain 2hsp 
SOLUTION STRUCTURE OF THE SH3 DOMAIN OF PHOSPHOLIPASE CGAMMA 2i0n 
Structure of Dictyostelium discoideum Myosin VII SH3 domain with adjacent proline rich region 2iim 
SH3 Domain of Human Lck 2j05 
Crystal structure of the RasGAP SH3 domain at 1.5 Angstrom resolution 2j06 
Crystal structure of the RasGAP SH3 domain at 1.8 Angstrom resolution 2j6f 
N-TERMINAL SH3 DOMAIN OF CMS (CD2AP HUMAN HOMOLOG) BOUND TO CBL-B PEPTIDE 2j6k 
N-TERMINAL SH3 DOMAIN OF CMS (CD2AP HUMAN HOMOLOG) 2j6o 
ATYPICAL POLYPROLINE RECOGNITION BY THE CMS N-TERMINAL SH3 DOMAIN. CMS:CD2 HETEROTRIMER 2j7i 
ATYPICAL POLYPROLINE RECOGNITION BY THE CMS N-TERMINAL SH3 DOMAIN. CMS:CD2 HETERODIMER 2jm8 
R21A Spc-SH3 free 2jm9 
R21A Spc-SH3 bound 2jma 
R21A Spc-SH3:P41 complex 2jmc 
Chimer between Spc-SH3 and P41 2js0 
Solution structure of second SH3 domain of adaptor Nck 2js2 
Solution structure of first SH3 domain of adaptor Nck 2jt4 
Solution Structure of the Sla1 SH3-3-Ubiquitin Complex 2jte 
Third SH3 domain of CD2AP 2jw4 
NMR solution structure of the N-terminal SH3 domain of human Nckalpha 2jxb 
Structure of CD3epsilon-Nck2 first SH3 domain complex 2k2m 
Structural Basis of PxxDY Motif Recognition in SH3 Binding 2k3b 
Seeing the Invisible: Structures of Excited Protein States by Relaxation Dispersion NMR 2k6d 
CIN85 Sh3-C domain in complex with ubiquitin 2k79 
Solution Structure of the binary complex between the SH3 and SH2 domain of interleukin-2 tyrosine kinase 2k7a 
Ensemble Structures of the binary complex between the SH3 and SH2 domain of interleukin-2 tyrosine kinase. 2k9g 
Solution structure of the third SH3 domain of the Cin85 adapter protein 2kbt 
Attachment of an NMR-invisible solubility enhancement tag (INSET) using a sortase-mediated protein ligation method 2ke9 
NMR solution structure of the CASKIN SH3 domain 2kea 
NMR solution structure of the HACS1 SH3 domain 2kgt 
Solution structure of SH3 domain of PTK6 2knb 
Solution NMR structure of the parkin Ubl domain in complex with the endophilin-A1 SH3 domain 2kr3 
Solution structure of SHA-D 2krm 
RDC refined solution structure of the first SH3 domain of CD2AP 2krn 
High resolution structure of the second SH3 domain of CD2AP 2kro 
RDC refined high resolution structure of the third SH3 domain of CD2AP 2krs 
Solution NMR structure of SH3 domain from CPF_0587 (fragment 415-479) from Clostridium perfringens. Northeast Structural Genomics Consortium (NESG) Target CpR74A. 2kt1 
Solution NMR Structure of the SH3 Domain from the p85beta subunit of Phosphatidylinositol 3-kinase from H.sapiens, Northeast Structural Genomics Consortium Target HR5531E 2kxc 
1H, 13C, and 15N Chemical Shift Assignments for IRTKS-SH3 and EspFu-R47 complex 2kxd 
The structure of SH3-F2 2kym 
Solution structure of the Bem1p SH3-CI domain from L.elongisporus in complex with Ste20p peptide 2l0a 
Solution NMR Structure of Signal transducing adapter molecule 1 STAM-1 from Homo sapiens, Northeast Structural Genomics Consortium Target HR4479E 2l2p 
Folding Intermediate of the Fyn SH3 A39V/N53P/V55L from NMR Relaxation Dispersion Experiments 2l3p 
Structure of the prolyl cis isomer of the Crk Protein 2l3q 
Structure of the prolyl trans isomer of the Crk Protein 2l3s 
Structure of the autoinhibited Crk 2lcs 
Yeast Nbp2p SH3 domain in complex with a peptide from Ste20p 2lj0 
The third SH3 domain of R85FL 2lj1 
The third SH3 domain of R85FL with ataxin-7 PRR 2lj3 
PFBD: High-throughput Strategy of Backbone fold Determination for small well-folded proteins in less than a day 2lmj 
Itk-sh3 2lnh 
Enterohaemorrhagic E. coli (EHEC) exploits a tryptophan switch to hijack host F-actin assembly 2lp5 
Native Structure of the Fyn SH3 A39V/N53P/V55L 2lqn 
Solution structure of CRKL 2lqw 
Solution structure of phosphorylated CRKL 2lx7 
Solution NMR structure of SH3 domain of growth arrest-specific protein 7 (GAS7) (fragment 1-60) from Homo sapiens, Northeast Structural Genomics Consortium (NESG) Target HR8574A 2lz6 
Distinct ubiquitin binding modes exhibited by sh3 domains: molecular determinants and functional implications 2m0y 
Solution structure of the SH3 domain of DOCK180 2m51 
NMR structure of the SH3 domain of human RAS p21 protein activator (GTPase activating protein) 1 2mcn 
Distinct ubiquitin binding modes exhibited by SH3 domains: molecular determinants and functional implications 2mio 
Solution NMR Structure of SH3 Domain 1 of Rho GTPase-activating Protein 10 from Homo sapiens, Northeast Structural Genomics Consortium (NESG) Target HR9129A 2mox 
2MOX 2nuz 
crystal structure of alpha spectrin SH3 domain measured at room temperature 2nwm 
Solution structure of the first SH3 domain of human Vinexin and its interaction with the peptides from Vinculin 2o2o 
Solution structure of domain B from human CIN85 PROTEIN 2o2w 
Extending powder diffraction to proteins: structure solution of the second SH3 domain from ponsin 2o31 
Crystal structure of the second SH3 domain from ponsin 2o88 
Crystal structure of the N114A mutant of ABL-SH3 domain complexed with a designed high-affinity peptide ligand: implications for SH3-ligand interactions 2o9s 
The second SH3 domain from ponsin 2o9v 
The second SH3 domain from Ponsin in complex with the paxillin proline rich region 2oaw 
Structure of SHH variant of "Bergerac" chimera of spectrin SH3 2oi3 
NMR Structure Analysis of the Hematopoetic Cell Kinase SH3 Domain complexed with an artificial high affinity ligand (PD1) 2oj2 
NMR Structure Analysis of the Hematopoetic Cell Kinase SH3 Domain complexed with an artificial high affinity ligand (PD1) 2p4r 
Structural basis for a novel interaction between AIP4 and beta-PIX 2pni 
SOLUTION STRUCTURE AND LIGAND-BINDING SITE OF THE SH3 DOMAIN OF THE P85ALPHA SUBUNIT OF PHOSPHATIDYLINOSITOL 3-KINASE 2ptk 
CHICKEN SRC TYROSINE KINASE 2pz1 
Crystal Structure of Auto-inhibited Asef 2rf0 
Crystal structure of human mixed lineage kinase MAP3K10 SH3 domain 2rmo 
Solution structure of alpha-spectrin_SH3-bergerac from Chicken 2rn8 
NMR structure note: murine Itk SH3 domain 2rna 
Itk SH3 average minimized 2rol 
Structural Basis of PxxDY motif recognition in SH3 binding 2rot 
Structure of chimeric variant of SH3 domain- SHH 2rpn 
A crucial role for high intrinsic specificity in the function of yeast SH3 domains 2rqr 
The solution structure of human DOCK2 SH3 domain - ELMO1 peptide chimera complex 2rqt 
Solution structure of the human DDEF1 SH3 domain 2rqu 
Solution structure of the complex between the DDEF1 SH3 domain and the APC SAMP1 motif 2rqv 
Solution structure of SH3CI domain of Bem1p 2rqw 
Solution structure of Bem1p SH3CI domain complexed with Ste20p-PRR peptide 2sem 
SEM5 SH3 DOMAIN COMPLEXED WITH PEPTOID INHIBITOR 2src 
CRYSTAL STRUCTURE OF HUMAN TYROSINE-PROTEIN KINASE C-SRC, IN COMPLEX WITH AMP-PNP 2v1q 
Atomic-resolution structure of the yeast Sla1 SH3 domain 3 2v1r 
Yeast Pex13 SH3 domain complexed with a peptide from Pex14 at 2.1 A resolution 2vge 
Crystal structure of the C-terminal region of human iASPP 2vkn 
YEAST SHO1 SH3 DOMAIN COMPLEXED WITH A PEPTIDE FROM PBS2 2vvk 
Grb2 SH3C (1) 2vwf 
Grb2 SH3C (2) 2w0z 
Grb2 SH3C (3) 2w10 
Mona SH3C in complex 2x3w 
structure of mouse syndapin I (crystal form 2) 2x3x 
structure of mouse syndapin I (crystal form 1) 2xkx 
Single particle analysis of PSD-95 in negative stain 2xmf 
Myosin 1e SH3 2ydl 
Crystal structure of SH3C from CIN85 2ysq 
Solution structure of the SH3 domain from Rho guanine nucleotide exchange factor 9 2yt6 
Solution structure of the SH3_1 domain of Yamaguchi sarcoma viral (v-yes) oncogene homolog 1 2yun 
Solution structure of the SH3 domain of human Nostrin 2yuo 
Solution structure of the SH3 domain of mouse RUN and TBC1 domain containing 3 2yup 
Solution structure of the second SH3 domain of human Vinexin 2yuq 
Solution structure of the SH3 domain of human Tyrosine-protein kinase ITK/TSK 3a98 
Crystal structure of the complex of the interacting regions of DOCK2 and ELMO1 3c0c 
X-ray Crystal Structure of the Rat Endophilin A2 SH3 Domain 3cqt 
N53I V55L MUTANT of FYN SH3 DOMAIN 3eg0 
Crystal structure of the N114T mutant of ABL-SH3 domain 3eg1 
Crystal structure of the N114Q mutant of ABL-SH3 domain complexed with a designed high-affinity peptide ligand: implications for SH3-ligand interactions 3eg2 
Crystal structure of the N114Q mutant of ABL-SH3 domain 3eg3 
Crystal structure of the N114A mutant of ABL-SH3 domain 3egu 
Crystal structure of the N114A mutant of ABL-SH3 domain 3ehq 
Crystal Structure of Human Osteoclast Stimulating Factor 3ehr 
Crystal Structure of Human Osteoclast Stimulating Factor 3fj5 
Crystal structure of the c-src-SH3 domain 3gbq 
SOLUTION NMR STRUCTURE OF THE GRB2 N-TERMINAL SH3 DOMAIN COMPLEXED WITH A TEN-RESIDUE PEPTIDE DERIVED FROM SOS DIRECT REFINEMENT AGAINST NOES, J-COUPLINGS, AND 1H AND 13C CHEMICAL SHIFTS, MINIMIZED AVERAGE STRUCTURE 3gf9 
Crystal structure of human Intersectin 2 RhoGEF domain 3h0f 
Crystal structure of the human Fyn SH3 R96W mutant 3h0h 
human Fyn SH3 domain R96I mutant, crystal form I 3h0i 
human Fyn SH3 domain R96I mutant, crystal form II 3haj 
Crystal structure of human PACSIN2 F-BAR domain (p212121 lattice) 3i35 
Human SH3 domain of protein LASP1 3i5r 
PI3K SH3 domain in complex with a peptide ligand 3i5s 
Crystal structure of PI3K SH3 3i9q 
Crystal Structure of the triple mutant S19G-P20D-R21S of alpha spectrin SH3 domain 3iql 
Crystal structure of the rat endophilin-A1 SH3 domain 3jbr 
3JBR 3jv3 
Structure of SH3E-DH unit of murine intersectin-1L 3kfv 
Crystal structure of the SH3-kinase fragment of tight junction protein 3 (TJP3) in apo-form 3lh5 
Crystal Structure of the SH3-Guanylate kinase core domain of ZO-1 3m0p 
Crystal Structure of the R21D mutant of alpha-spectrin SH3 domain. Crystal obtained in ammonium sulphate at pH 4. 3m0q 
Crystal Structure of the R21D mutant of alpha-spectrin SH3 domain. Crystal obtained in ammonium sulphate at pH 5. 3m0r 
Crystal Structure of the R21D mutant of alpha-spectrin SH3 domain. Crystal obtained in ammonium sulphate at pH 6. 3m0s 
Crystal Structure of the R21D mutant of alpha-spectrin SH3 domain. Crystal obtained in ammonium sulphate at pH 7 3m0t 
Crystal Structure of the R21D mutant of alpha-spectrin SH3 domain. Crystal obtained in ammonium sulphate at pH 9. 3m0u 
Crystal Structure of the R21D mutant of alpha-spectrin SH3 domain. Hexagonal crystal obtained in sodium formate at pH 6.5. 3ngp 
High resolution structure of alpha-spectrin SH3 domain mutant with a redesigned core 3nhn 
Crystal structure of the SRC-family kinase HCK SH3-SH2-linker regulatory region 3nmz 
Crytal structure of APC complexed with Asef 3o5z 
Crystal structure of the SH3 domain from p85beta subunit of phosphoinositide 3-kinase (PI3K) 3pvl 
Structure of myosin VIIa MyTH4-FERM-SH3 in complex with the CEN1 of Sans 3qwx 
CED-2 1-174 3qwy 
CED-2 3rbb 
HIV-1 NEF protein in complex with engineered HCK SH3 domain 3rea 
HIV-1 Nef protein in complex with engineered Hck-SH3 domain 3reb 
HIV-1 Nef protein in complex with engineered Hck-SH3 domain 3rnj 
Crystal structure of the SH3 domain from IRSp53 (BAIAP2) 3sem 
SEM5 SH3 DOMAIN COMPLEXED WITH PEPTOID INHIBITOR 3shw 
Crystal structure of ZO-1 PDZ3-SH3-Guk supramodule complex with Connexin-45 peptide 3thk 
Structure of SH3 chimera with a type II ligand linked to the chain C-terminal 3tsw 
crystal structure of the PDZ3-SH3-GUK core module of Human ZO-1 3tsz 
crystal structure of PDZ3-SH3-GUK core module from human ZO-1 in complex with 12mer peptide from human JAM-A cytoplasmic tail 3tvt 
Structural basis for Discs Large interaction with Pins 3u23 
Atomic resolution crystal structure of the 2nd SH3 domain from human CD2AP (CMS) in complex with a proline-rich peptide from human RIN3 3ua6 
Crystal Structure of the Human Fyn SH3 domain 3ua7 
Crystal Structure of the Human Fyn SH3 domain in complex with a peptide from the Hepatitis C virus NS5A-protein 3uat 
Guanylate Kinase Domains of the MAGUK Family Scaffold Proteins as Specific Phospho-Protein Binding Modules 3uf4 
Crystal structure of a SH3 and SH2 domains of FYN protein (Proto-concogene Tyrosine-protein kinase Fyn) from Mus musculus at 1.98 A resolution 3ulr 
Lysozyme contamination facilitates crystallization of a hetero-trimericCortactin:Arg:Lysozyme complex 3vry 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 4-Amino-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-7-yl-cyclopentane 3vrz 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 1-[4-(4-amino-7-cyclopentyl-7H-pyrrolo[2,3-d]pyrimidin-5-yl)phenyl]-3-benzylurea 3vs0 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor N-[4-(4-amino-7-cyclopentyl-7H-pyrrolo[2,3-d]pyrimidin-5-yl)phenyl]benzamide 3vs1 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 1-[4-(4-amino-7-cyclopentyl-7H-pyrrolo[2,3-d]pyrimidin-5-yl)phenyl]-3-phenylurea 3vs2 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 7-[cis-4-(4-methylpiperazin-1-yl)cyclohexyl]-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine 3vs3 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 7-[trans-4-(4-methylpiperazin-1-yl)cyclohexyl]-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine 3vs4 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 5-(4-phenoxyphenyl)-7-(tetrahydro-2H-pyran-4-yl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine 3vs5 
Crystal structure of HCK complexed with a pyrrolo-pyrimidine inhibitor 7-(1-methylpiperidin-4-yl)-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine 3vs6 
Crystal structure of HCK complexed with a pyrazolo-pyrimidine inhibitor tert-butyl {4-[4-amino-1-(propan-2-yl)-1H-pyrazolo[3,4-d]pyrimidin-3-yl]-2-methoxyphenyl}carbamate 3vs7 
Crystal structure of HCK complexed with a pyrazolo-pyrimidine inhibitor 1-cyclopentyl-3-(1H-pyrrolo[2,3-b]pyridin-5-yl)-1H-pyrazolo[3,4-d]pyrimidin-4-amine 3zl7 
BACE2 FYNOMER COMPLEX 4a63 
Crystal structure of the p73-ASPP2 complex at 2.6A resolution 4afq 
Human Chymase - Fynomer Complex 4afs 
Human Chymase - Fynomer Complex 4afu 
Human Chymase - Fynomer Complex 4afz 
Human Chymase - Fynomer Complex 4ag1 
Human Chymase - Fynomer Complex 4ag2 
Human Chymase - Fynomer Complex 4bne 
Pacsin2 Interacts with Membranes and Actin-Filaments 4cc2 
Complex of human Tuba C-terminal SH3 domain with human N-WASP proline- rich peptide - P212121 4cc3 
Complex of human Tuba C-terminal SH3 domain and Mena proline-rich peptide - H3 4cc4 
Complex of InlC of Listeria monocytogenes and human Tuba C-terminal SH3 domain 4cc7 
Crystal structure of the sixth or C-terminal SH3 domain of human Tuba in complex with proline-rich peptides of N-WASP. Space group P41 4d8d 
Crystal structure of HIV-1 NEF Fyn-SH3 R96W variant 4d8k 
Crystal structure of a SH3-SH2 domains of a lymphocyte-specific protein tyrosine kinase (LCK) from Homo sapiens at 2.36 A resolution 4dex 
Crystal structure of the Voltage Dependent Calcium Channel beta-2 Subunit in Complex With The CaV2.2 I-II Linker. 4dey 
Crystal structure of the Voltage Dependent Calcium Channel beta-2 Subunit in Complex With The CaV1.2 I-II Linker. 4e6r 
Crystal structure of a Cytoplasmic protein NCK2 (NCK2) from Homo sapiens at 2.20 A resolution 4eik 
Crystal Structure of the Human Fyn SH3 domain in complex with the synthetic peptide VSL12 4esr 
Molecular and Structural Characterization of the SH3 Domain of AHI-1 in Regulation of Cellular Resistance of BCR-ABL+ Chronic Myeloid Leukemia Cells to Tyrosine Kinase Inhibitors 4f14 
Structure of the SH3 domain of human nebulette in complex with a peptide of XIRP2 4f16 
Crystal Structure of the alpha spectrin SH3 domain at pH 5 4f17 
Crystal Structure of the alpha spectrin SH3 domain at pH 9 4fss 
Crystal structure of a RAS p21 protein activator (RASA1) from Homo sapiens at 2.25 A resolution 4gbq 
SOLUTION NMR STRUCTURE OF THE GRB2 N-TERMINAL SH3 DOMAIN COMPLEXED WITH A TEN-RESIDUE PEPTIDE DERIVED FROM SOS DIRECT REFINEMENT AGAINST NOES, J-COUPLINGS, AND 1H AND 13C CHEMICAL SHIFTS, 15 STRUCTURES 4glm 
Crystal structure of the SH3 Domain of DNMBP protein [Homo sapiens] 4hck 
HUMAN HCK SH3 DOMAIN, NMR, 25 STRUCTURES 4hvu 
Crystal structure of the T98D c-Src-SH3 domain mutant in complex with the high affinity peptide APP12 4hvv 
Crystal structure of the T98E c-Src-SH3 domain mutant in complex with the high affinity peptide APP12 4hvw 
Crystal structure of the T98E c-Src-SH3 domain mutant in complex with the high affinity peptide VSL12 4hxj 
Crystal structure of SH3:RGT complex 4hzs 
Crystal structure of Ack1 kinase domain with C-terminal SH3 domain 4igz 
Crystal structure of the SH3 domain of human sorbin and SH3 domain-containing protein 2 4iim 
Crystal structure of the Second SH3 Domain of ITSN1 bound with a synthetic peptide 4iio 
Crystal Structure of the Second SH3 Domain of ITSN2 Bound with a Synthetic Peptide 4j9b 
Crystal structure of the Abl-SH3 domain H59Q-N96T mutant 4j9c 
Crystal structure of the Abl-SH3 domain H59Q-N96T mutant complexed with the designed high-affinity peptide ligand P17 4j9d 
Crystal structure of the N114A mutant of the Abl-SH3 domain complexed with the high affinity peptide P0 4j9e 
Crystal structure of the N114A mutant of the Abl-SH3 domain complexed with the high affinity peptide P17 4j9f 
Crystal structure of the Abl-SH3 domain complexed with the high affinity peptide P0 4j9g 
Crystal structure of the ABL-SH3 domain complexed with the designed high-affinity peptide ligand P7 at pH7 4j9h 
Crystal structure of the ABL-SH3 domain complexed with the designed high-affinity peptide ligand P7 at pH 8 4j9i 
Crystal structure of the ABL-SH3 domain complexed with the designed high-affinity peptide ligand P17 4jjb 
Crystal structure of the Abl-SH3 domain at pH3 4jjc 
Crystal structure of the Abl-SH3 domain at pH5 4jjd 
Crystal structure of the N114A Abl-SH3 domain mutant at pH4 4jz3 
Crystal structure of the chicken c-Src-SH3 domain intertwined dimer 4jz4 
Crystal structure of chicken c-Src-SH3 domain: monomeric form 4k11 
The structure of 1NA in complex with Src T338G 4le9 
Crystal structure of a chimeric c-Src-SH3 domain 4ln2 
4LN2 4lnp 
4LNP 4lud 
Crystal Structure of HCK in complex with the fluorescent compound SKF86002 4lue 
Crystal Structure of HCK in complex with 7-[trans-4-(4-methylpiperazin-1-yl)cyclohexyl]-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine (resulting from displacement of SKF86002) 4m4z 
SH3 and SH2 domains of human Src-like adaptor protein 2 (SLAP2) 4mt6 
4MT6 4mt7 
4MT7 4n78 
The WAVE Regulatory Complex Links Diverse Receptors to the Actin Cytoskeleton 4oml 
4OML 4omm 
4OMM 4omn 
4OMN 4omo 
4OMO 4omp 
4OMP 4omq 
4OMQ 4orz 
HIV-1 Nef protein in complex with single domain antibody sdAb19 and an engineered Hck SH3 domain 4qt7 
4QT7 4rtt 
4RTT 4rtu 
4RTU 4rtv 
4RTV 4rtw 
4RTW 4rtx 
4RTX 4rty 
4RTY 4rtz 
4RTZ 4rug 
4RUG 4u5w 
4U5W 4wci 
4WCI 4wsi 
4WSI 4x1v 
4X1V 4xi2 
4XI2 4y92 
4Y92 4z88 
4Z88 4z89 
4Z89 4z8a 
4Z8A 4znx 
4ZNX 4zw2 
4ZW2 5ec7 
5EC7 5eca 
5ECA 5f3y 
5F3Y 5ful 
5FUL 5fwa 
5FWA 5gjv 
5GJV 5gjw 
5GJW 5hck 
HUMAN HCK SH3 DOMAIN, NMR, MINIMIZED AVERAGE STRUCTURE 5i11 
5I11 5i22 
5I22 5ih2 
5IH2 5ixb 
5IXB 5l23 
5L23 - Links (links to other resources describing this domain)
-
INTERPRO IPR001452 PFAM SH3 PROSITE SH3

